Tracks
This chapter explains how to visualize tracks, how to retrieve reference data and finally how to perform generic comparisons between tracks.
A track is the fundamental building block for NGS analysis in the CLC Genomics Workbench. The idea behind tracks is to provide a unified framework for the visualization, comparison and analysis of genome-scale studies such as whole-genome sequencing or exome resequencing projects and a variety of different -Seq data (i.e. RNA-seq, ChIP-Seq, DNAse-Seq).
In tracks, all information is tied to genomic positions. A central coordinate-system is provided by a reference genome, which allows different types of data or results for different samples to be seen and analyzed together.
Different types of data are represented in different types of tracks, and each type of track has its own particular editors. An example of a paired-end mapping read-track displaying reads and coverage is shown in figure 23.1.
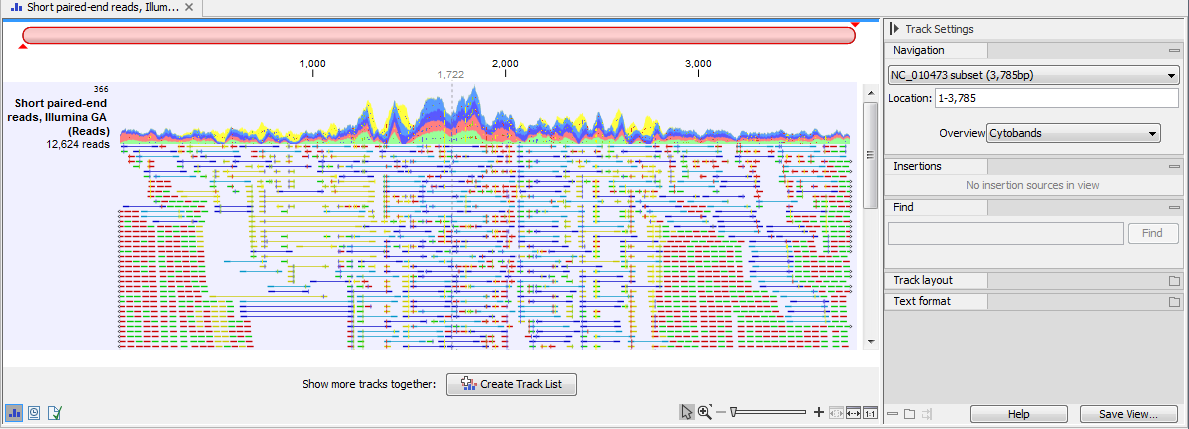
Figure 23.1: A paired-end mapping read-track opened, displaying reads and coverage. Below the track, the button for creating a Track List is visible. On the right is the Side Panel.
Subsections
- Track types
- Track lists
- Retrieving reference data tracks
- Merge Annotation Tracks
- Track Conversion
- Annotate and Filter
- Graphs
