I want to BLAST against my own sequence database, is this possible?
It is possible to download the entire BLAST program package and use it on your own computer, institution computer cluster or similar. This is preferred if you want to search in proprietary sequences or sequences unavailable in the public databases stored at NCBI. The downloadable BLAST package can either be installed as a web-based tool or as a command line tool. It is available for a wide range of different operating systems.The BLAST package can be downloaded free of charge from the following location http://www.ncbi.nlm.nih.gov/BLAST/download.shtml
Pre-formatted databases are available from a dedicated BLAST ftp site ftp://ftp.ncbi.nlm.nih.gov/blast/db/. Moreover, it is possible to download programs/scripts from the same site enabling automatic download of changed BLAST databases. Thus it is possible to schedule a nightly update of changed databases and have the updated BLAST database stored locally or on a shared network drive at all times. Most BLAST databases on the NCBI site are updated on a daily basis to include all recent sequence submissions to GenBank.
A few commercial software packages are available for searching your own data. The advantage of using a commercial program is obvious when BLAST is integrated with the existing tools of these programs. Furthermore, they let you perform BLAST searches and retain annotations on the query sequence (see figure 12.24). It is also much easier to batch download a selection of hit sequences for further inspection.
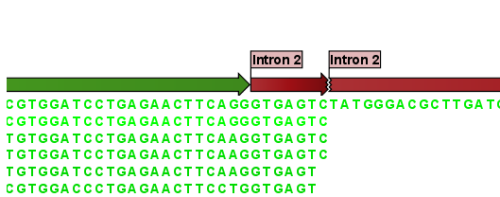
Figure 12.24: Snippet of alignment view of BLAST results. Individual alignments are represented directly in a graphical view. The top sequence is the query sequence and is shown with a selection of annotations.
