Identify Differentially Expressed Gene Groups and Pathways
This tool can be used to investigate candidate differentially expressed genes
for a common functional role.
For example if you would like to compare different cancer patients to check
whether e.g. the same pathways are affected in different individuals, you can use this tool.
For this, you need a GO association file, which includes gene names and associated Gene Ontology terms.
A GO association file with the top-level GO terms annotated (GO slim) is
provided with the CLC Genomics Workbench and can be downloaded using the
Data Management (![]() ) function found in the top right
corner of the Workbench (see Download and configure reference data).
) function found in the top right
corner of the Workbench (see Download and configure reference data).
To run the analysis go to the toolbox:
Legacy tools
| Identify Differentially Expressed Gene Groups and
Pathways (![]() )
)
When you run the Identify Differentially Expressed Gene Groups and Pathways
analysis, you first have to select the expression comparison track
(![]() ) you wish to annotate with the GO term enrichment analysis.
Expression comparison tracks can be created e.g. by the create fold change track
tool (see Create fold
change track).
) you wish to annotate with the GO term enrichment analysis.
Expression comparison tracks can be created e.g. by the create fold change track
tool (see Create fold
change track).
After clicking Next, you have to specify the annotation association file, a gene track, and finally which ontology (cellular component, biological process or molecular function) you would like to test for (see figure 33.6).
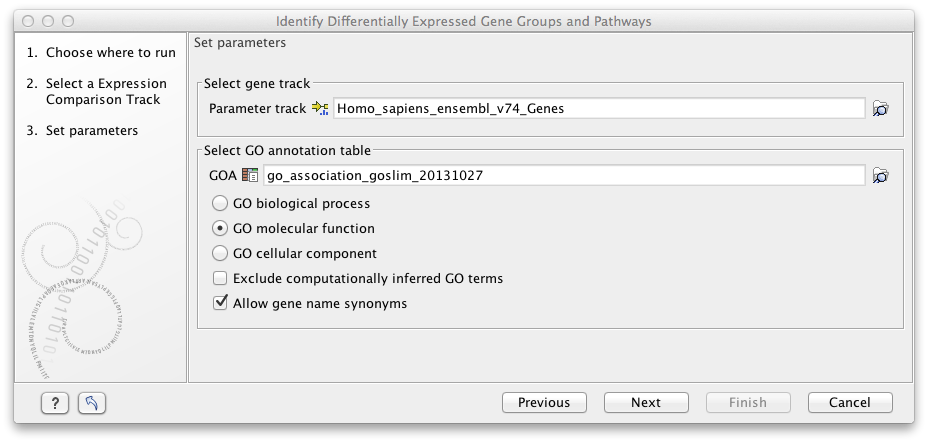
Figure 33.6:
Select gene track, GO annotation table, and ontology.
Next, the Workbench tries to match gene names from the expression comparison track with the gene names in the GO association file. Please be aware that the same gene name definition should be used in both files.
Based on this, the Workbench finds GO terms that are over-represented in the list. A hypergeometric test is used to identify over-represented GO terms by testing whether some of the GO terms are over-represented in a given gene set, compared to a randomly selected set of genes.
The result is a table with GO terms and the calculated p-value for the differentially expressed genes, and a new expression comparison track with annotated GO terms and the corresponding p-value (see figure 33.7). The p-value is the probability of obtaining a test statistic at least as extreme as the one that was actually observed, or in other words how significant (trustworthy) a result is. In case of a small p-value the probability of achieving the same result by chance with the same test statistic is very small.
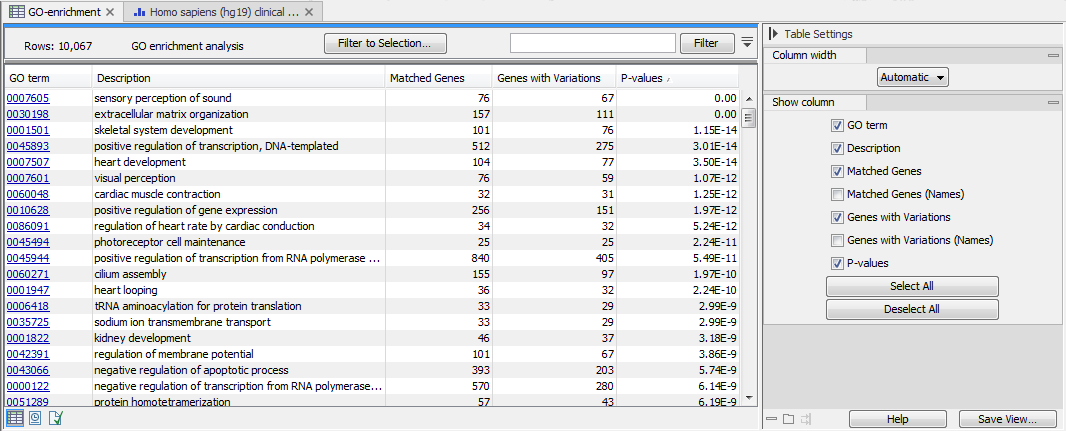
Figure 33.7: The results of the analysis.
