Filter Variants on Custom Criteria
The Filter Variants on Custom Criteria tool can be used to identify and extract variants that fulfill certain criteria. This tool provides the exact same functionality than the one offered by filtering a variant table. There is thus no advantage in using this tool on its own as filtering a table is usually easier, but the tool can be used when building workflows where a filtering step is needed, or when working in batch mode.
To run the Filter Variants on Custom Criteria, go to:
Toolbox | Resequencing Analysis | Variant Filtering | Filter Variants on Custom Criteria (![]() )
)
In the first step, select a variant track as input and click Next.
In the Filter criteria dialog, specify a variant track that contains at least all the criteria you want to filter the first variant track on by using the browse icon (![]() ). Once specified, click on the Load Annotations button to create the drop down menu available in the filter field below. Note that in this context, we call this track a guidance track because it will define what are the filter criteria available; in the dialog, Load Annotations will transform every column headers of the guidance variant track in a filtering criteria. Please note, that when you create filter criteria with a guiding variant track, you can only filter on annotations that are present in the variant tracks that you would like to filter.
). Once specified, click on the Load Annotations button to create the drop down menu available in the filter field below. Note that in this context, we call this track a guidance track because it will define what are the filter criteria available; in the dialog, Load Annotations will transform every column headers of the guidance variant track in a filtering criteria. Please note, that when you create filter criteria with a guiding variant track, you can only filter on annotations that are present in the variant tracks that you would like to filter.
An example of filter criteria is shown in figure 27.24. In this example, we filter away all variants that are not found on chromosome 1 and that are not homozygous. As a result you will only keep the variants that are homozygous and found on chromosome 1. The bottom filter field on the figure shows the various other filter criteria available from the guidance variant track.
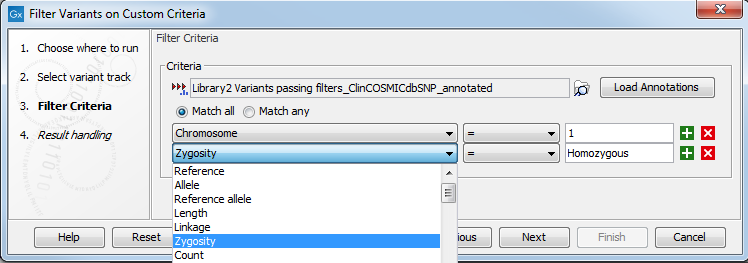
Figure 27.24: Create filter criteria to extract the variants of interest. In this example we will extract the homozygous variants found on chromosome 1.
The output from the Filter Variants on Custom Criteria tool is a variant track (and table) that contains only the variants that fulfill the specified filter criteria.
