Reads track
A track containing the mapped reads is generated. Details about viewing and editing of reads-tracks are described in tracks, and the resequencing.
If you have chosen the strand specific option when setting up your analysis, it may be helpful to note that the colors of mapped single reads represent the orientation of the read relative to the reference provided. When a gene track is provided along with the reference genome, the reads will be mapped using the strand you specified, but the coloring of the read will be relative to the reference gene. If the reads matches the orientation of the gene it is colored green, and if it is opposite to the orientation of the gene it is colored red. A summary list of the colors to expect with different combinations of gene orientation and strand specific mapping options is:
- Strand specific, forward orientation chosen + gene on plus strand of reference = single reads colored green.
- Strand specific, forward orientation chosen + gene on minus strand of reference = single reads colored red.
- Strand specific, reverse orientation chosen + gene on plus strand of reference = single reads colored red.
- Strand specific, reverse orientation chosen + gene on minus strand of reference = single reads colored green.
See figure 28.15 for an example of forward and reverse reads mapped to a gene on the plus strand.
Note: Reads mapping to intergenic regions will not be mapped in a strand specific way.
Although paired reads are coloured blue, they can be viewed as red and green 'single' reads by selecting the Disconnect paired reads box, within the Read Mapping Settings bar on the right-hand side of the track.
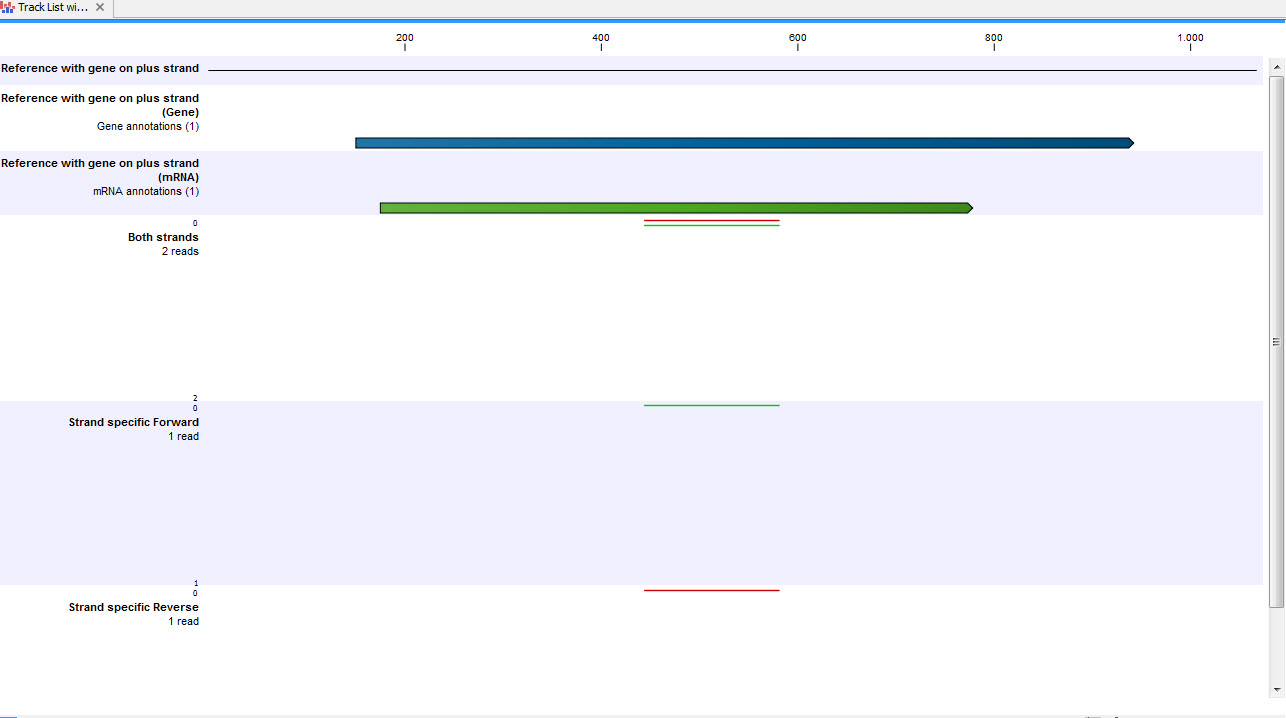
Figure 28.15: A track list showing a gene and transcript on the plus strand, and various mapping results. The first reads track shows a mapping of two reads (one 'forward' and one 'reverse') using strand specific 'both' option. Both reads map successfully; the forward read coloured green (because it matches the direction of the gene), and the reverse read coloured red. The second reads track shows a mapping of the same reads using strand specific ('forward') option. The reverse read does not map because it is not in the correct direction, therefore only the green forward read is shown. The final reads track shows a mapping of the same reads again but using strand specific 'reverse' option. This time, the green forward read does not map because it is in the wrong direction, and only the red reverse read is shown.
