Executing a workflow
Once installed and configured, a workflow will appear in the Toolbox under Installed Workflows (
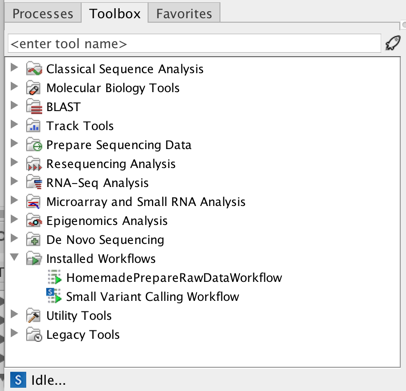
Figure 10.50: A workflow is installed and ready to be used.
The workflow is executed just as any other tool in the Toolbox by double-clicking or selecting it in the menu (or with the shortcut Ctrl + Enter). This will open a dialog where you select (and potentially download) relevant reference data, provide input data and configure the tools whose parameters have been left unlocked. Most workflows can also be run in batch mode (see Batch processing). In the last page of the dialog, you can preview all the parameters of the workflow, as well as the input data, before clicking "Next" to choose where to save the output, and then "Finish" to execute the workflow.
If you are connected to a CLC Genomics Server, you will be presented with the option to run the workflow locally on the Workbench or on the Server. When you are selecting where to run the workflow, you should also see a message should there be any missing configurations. There are more details about running Workflows here: http://resources.qiagenbioinformatics.com/manuals/clcgenomicsworkbench/current/index.php?manual=Workflows.html.
When the workflow is started, you can see the log file with detailed information from each step in the process.
If the workflow is not properly configured, you will see that in the dialog when the workflow is started 10.1.
Footnotes
- ... started10.1
- If the workflow uses a tool that is part of a plugin, a missing plugin can also be the reason why the workflow is not enabled. A workflow can also become outdated because the underlying tools have changed since the workflow was created (see Workflow identification and versioning)
