Type among Multiple Species (legacy)
Note this is the legacy version of the workflow found in section Type Among Multiple Species. We recommend avoiding using legacy workflows whenever possible. If you find missing functionality, please get in touch at ts-bioinformatics@qiagen.com.
The Type among Multiple Species (legacy) workflow is designed for typing a sample among multiple predefined species (figure 24.31).
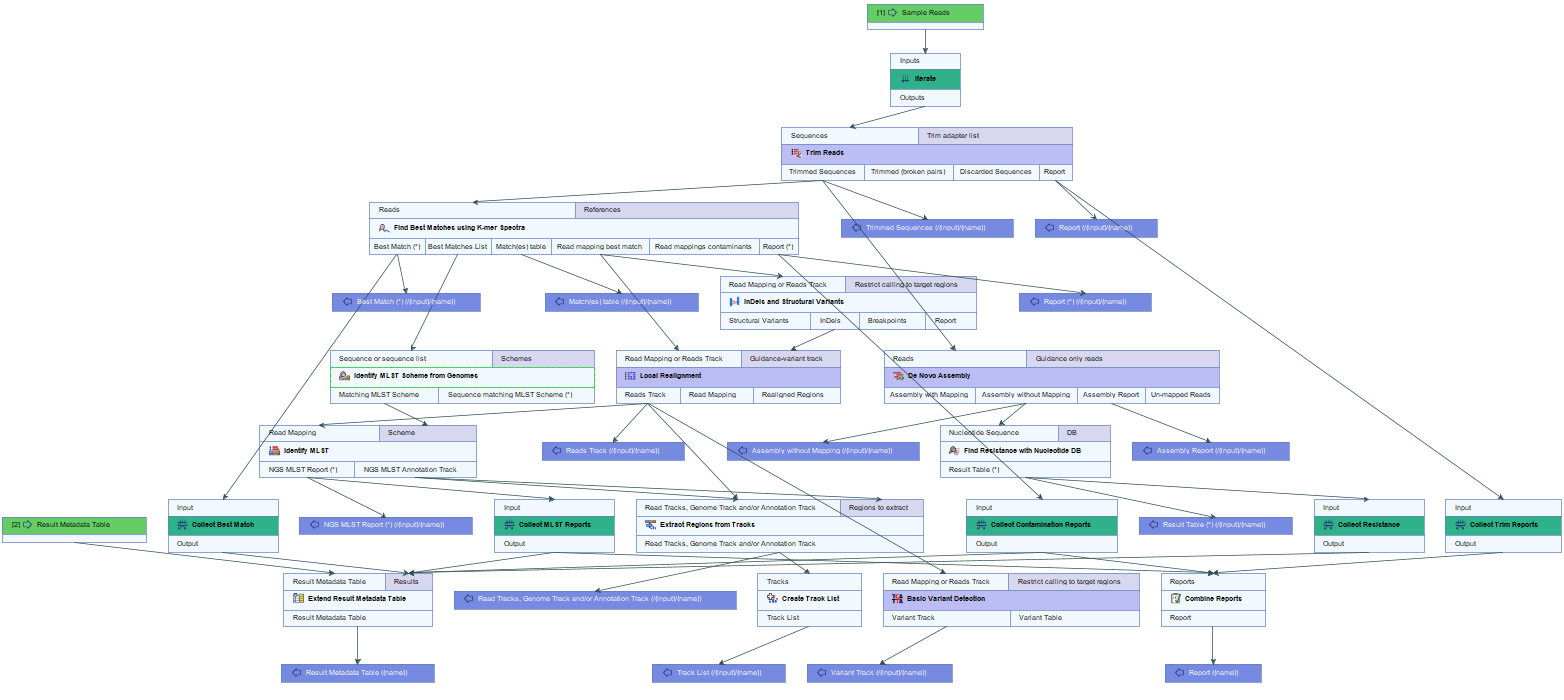
Figure 24.31: Overview of the template Type among Multiple Species (legacy) workflow.
Similar to the Type a Known Species (legacy) workflow, it allows identification of the closest matching reference species among the user specified reference list(s), but in this case, the list(s) may represent multiple species. The workflow identifies the associated MLST scheme and type, determines variants found when mapping the sample data against the identified best matching reference, and finds occurring resistance genes if they match genes within the user specified resistance database.
The workflow also automatically associates the analysis results to the user specified Result Metadata Table. For details about searching and quick filtering among the sample metadata and generated analysis result data(see Filtering in Result Metadata Table).
Subsections
- Preliminary steps to run the Type Multiple Species workflow
- How to run the Type among Multiple Species (legacy) workflow
- Example of results obtained using the Type among Multiple Species (legacy) workflow
