Example of results obtained using the Type a Known Species workflow
In this section, sequence data from five Salmonella enterica samples were typed using a customized Salmonella enterica version of the Type a Known Species workflow: the specified reference and the MLST scheme was specified as Salmonella enterica. The created Result Metadata Table shows these five samples and associated metadata for sample with accession no ERR277212.
Once the workflow analysis was performed, additional columns listing the analysis results (e.g., MLST, best matching reference and identified resistance genes) have automatically been added to the Result Metadata Table (figure 10.34). To ease the overview regarding applied analysis parameters, columns stating the workflow specified reference list, MLST scheme and resistance gene database have also been added automatically to the table.
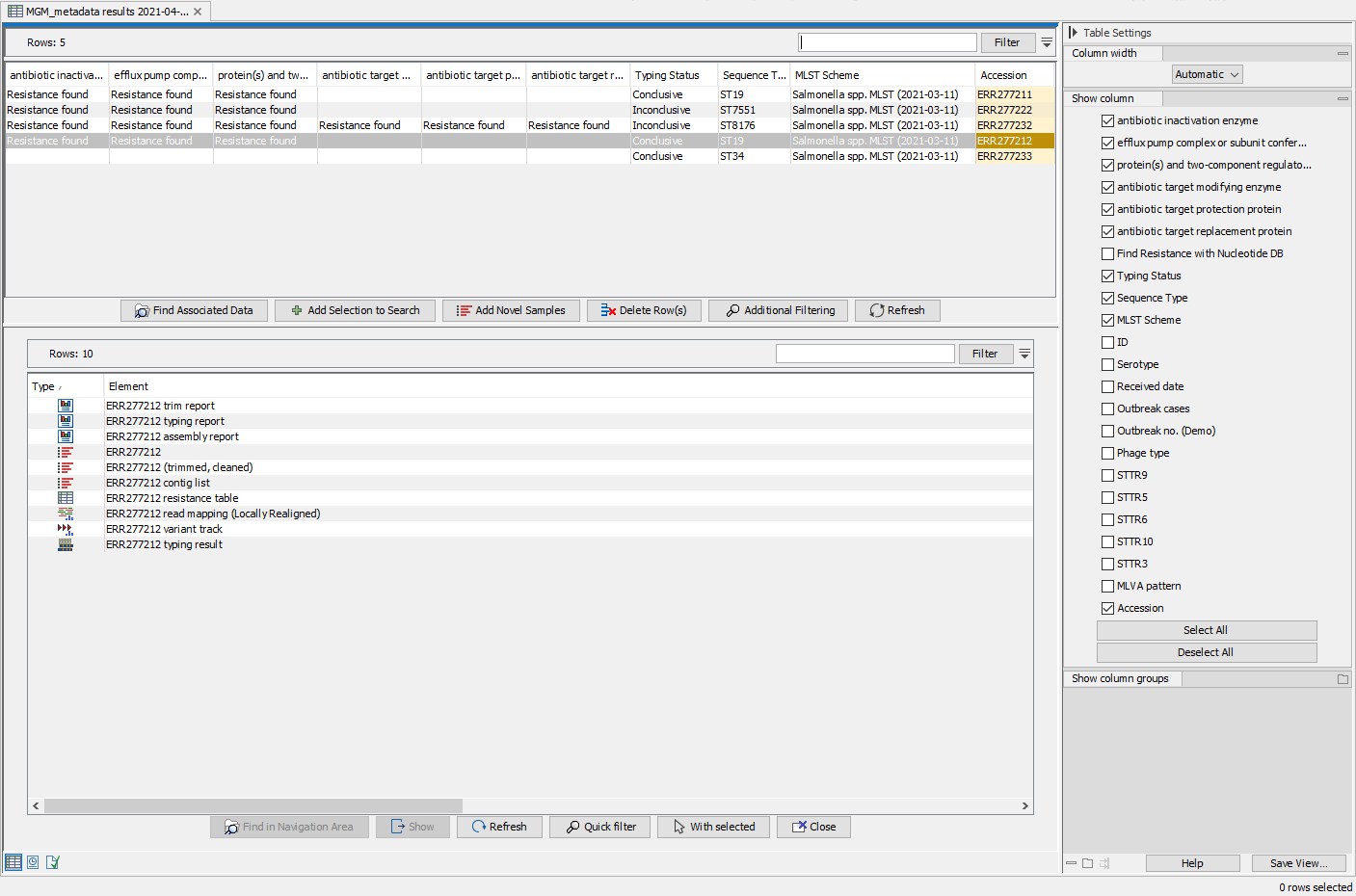
Figure 10.34: View of the updated Result Metadata Table and associated elements once the Type a Known Species workflow has been executed on the Salmonella (acc no ERR277212) sample.
Analyzing samples in batch will produce a large amount of output files, making it necessary to filter for the information you are looking for. Through the Result Metadata Table, it is possible to filter among sample metadata and analysis results. By clicking Find Associated Data (![]() ) and optionally performing additional filtering, it is possible to perform additional analyses on a selected subset directly from this Table, such as:
) and optionally performing additional filtering, it is possible to perform additional analyses on a selected subset directly from this Table, such as:
- Generation of SNP trees based on the same reference used for read mapping and variant detection (Create SNP Tree).
- Generation of K-mer Trees for identification of the closest common reference across samples (Create K-mer Tree).
- Run validated workflows (workflows that are associated with a Result Metadata Table and saved in your Navigation Area).
Note that the tool will output, among other files, variant tracks. It is possible to export multiple variant track files from monoploid data into a single VCF file with the Multi-VCF exporter. This exporter becomes available when installing the CLC Microbial Genomics Module. All variant track files must have the same reference genome for the Multi-VCF export to work.
