Example of results obtained using the Type among Multiple Species (legacy) workflow
The following example includes typing of 5 samples: 2 Salmonella enterica (acc nos ERR274480, ERR277203), 1 Listeria monocytogenes (acc no DRR000982) and 2 Shigella (acc nos ERR279274, ERR279296). Using the above customization of the Type among Multiple Species (legacy) workflow, analysis results are automatically summarized in the Result Metadata Table as shown in figure 24.43. The analysis results in this example include the name and the description of the best matching reference, the identified MLST and applied scheme, and some cases where aminoglycoside resistance was found. This is not the case in this example, but you could also choose (using options in the Table Setting window) to display additional information such as the 'applied best match' and 'resistance databases' for example.
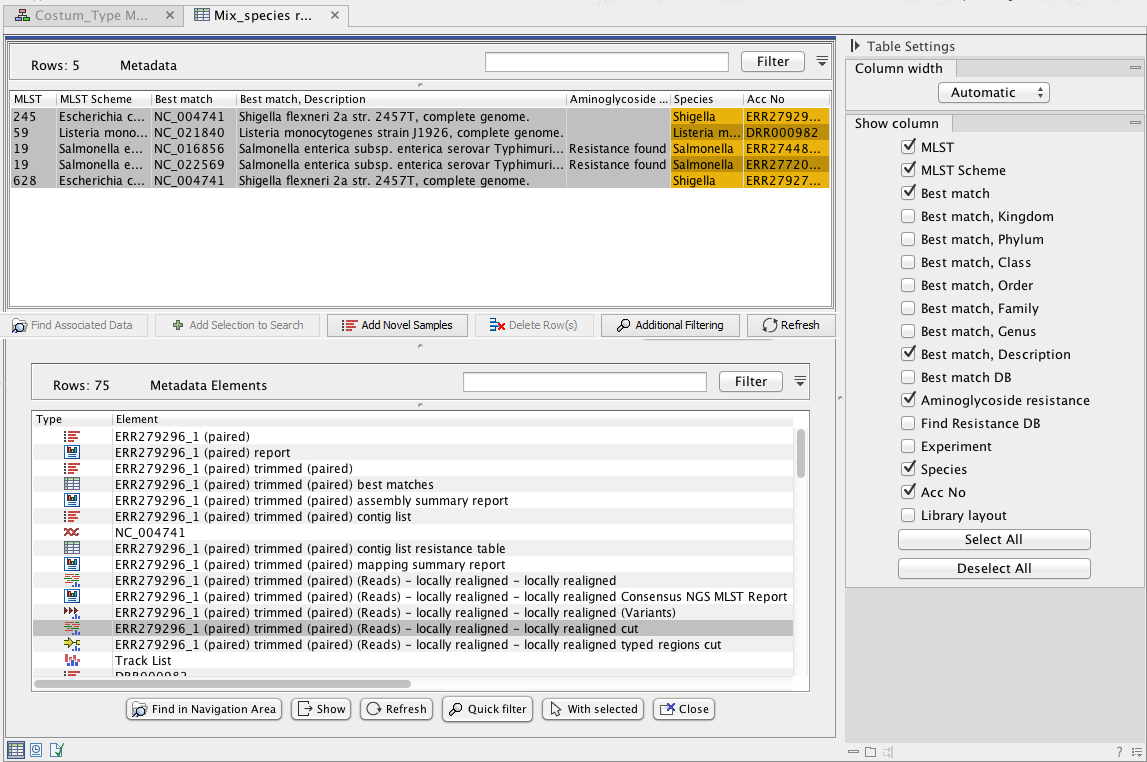
Figure 24.43: View of Result Metadata Table once the Type among Multiple Species (legacy) workflow has been executed (top) and associated data elements found (bottom).
By selecting samples in the Result Metadata Table, additional analyses can be performed directly from this Table:
- Generation of SNP trees based on the same reference used for read mapping and variant detection (Create SNP Tree).
- Generation of K-mer Trees for identification of the closest common reference across samples (Create K-mer Tree).
- Run validated workflows (workflows that are associated with a Result Metadata Table and saved in your Navigation Area).
Note that the tool will output, among other files, variant tracks. It is possible to export multiple variant track files from monoploid data into a single VCF file with the Multi-VCF exporter. This exporter is uploaded to the workbench when installing the Microbial Genomics Module. All variant track files must have the same reference genome for the Multi-VCF export to work.
