Data QC and Clean Host DNA
The Data QC and Clean Host DNA workflow performs trimming of reads, creates a QC report and cleans the dataset from host DNA, leaving back only the reads that do not match the host genome. To run the Data QC and Clean Host DNA workflow, go to Metagenomics (
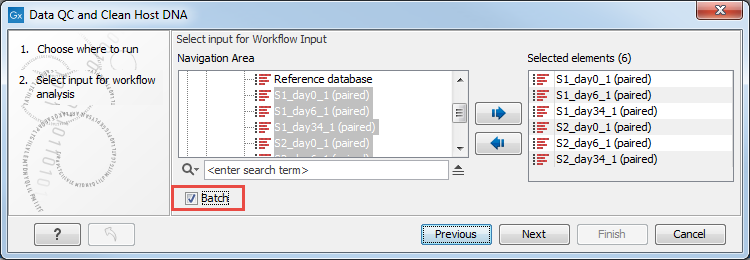
Figure 6.31: Select the reads.
In the "Trim Sequences" dialog, you can specify a trim adapter list and set up parameters if you would like to trim your sequences from adapters (figure 6.32).
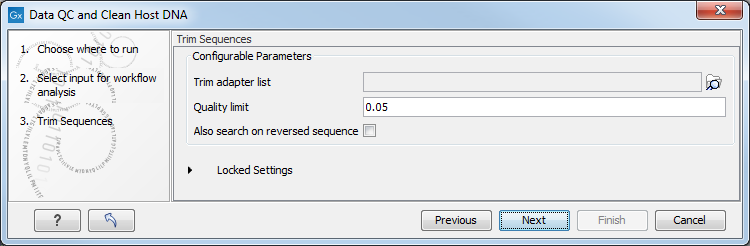
Figure 6.32: You can choose to trim adapter sequences from your sequencing reads.
The parameters that can be set are:
- Quality limit: defines the minimal value of the Phred score for which bases will not be trimmed.
- Also search on reversed sequence: the adapter sequences will also be searched on reverse sequences.
In the "Taxonomic Profiling" dialog, select the reference database index you will use to map the reads (figure 6.33). It is also possible to "Filter host reads". You must then specify the index of the host genome (in the case of human microbiota, the Homo sapiens GRCh38 for example). The reference database can be obtained by using the Download Curated Microbial Reference Database tool (Download Curated Microbial Reference Database) or Download Custom Microbial Reference Database tool (Download Custom Microbial Reference Database). The host index and (if using the custom downloader) the microbial reference index are built with the Create Taxonomic Profiling Index tool (Create Taxonomic Profiling Index).
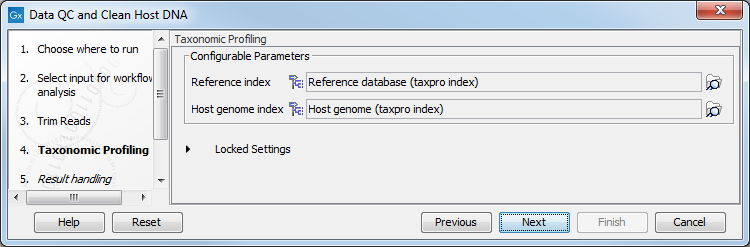
Figure 6.33: Select the reference databaes, and potentially choose to filter against an host genome to remove possible contamination.
The workflow will output a sequence list with reads cleaned from host DNA. In addition, it generates three reports: a trimming report, a graphical QC report and a supplementary QC report. All of these should be inspected in order to determine whether the quality of the sequencing reads and the trimming are acceptable. For a detailed description of the QC reports and indication on how to interpret the different values, see http://resources.qiagenbioinformatics.com/manuals/clcgenomicsworkbench/current/index.php?manual=QC_Sequencing_Reads.html.
For the trimming report, see http://resources.qiagenbioinformatics.com/manuals/clcgenomicsworkbench/current/index.php?manual=Trim_output.html.
