Batch launching workflows with multiple inputs
This section describes the launching of workflows with multiple inputs, where all input elements will be changed per batch. This launch mechanism is not intended for workflows with multiple input elements where one of the input elements remains the same in all batches, such as workflows meant to compare several tissues to a unique control tissue. At the moment, batch launching of such workflows is not possible, unless the common item is saved under different names as many times as there should be batches.
For workflows with multiple inputs where the inputs all need to change for each batch run, information specifying the grouping of the data elements and what role each element plays in a given analysis needs to be imported into the system from an Excel spreadsheet.
The requirements for launching such workflows in batch mode are:
- The workflow must be installed on the Workbench, meaning that the workflow is accessible from the Toolbox (as opposed to workflows accessible from the Navigation Area). See Distributing and installing workflows to learn how to install a workflow.
- The workflow is characterized by more than one input file, and all input elements are unique per batch. You cannot reuse a common input element (such as control reads for example), unless it has been saved under different names in the Navigation Area.
- An Excel format file (.xlsx/.xls) must be provided, with at least 3 different columns:
- Unique ID The first column must contain either the exact name of the data elements to be used as inputs, or partial name information such that data elements being entered into the analysis can be uniquely identified and matched with the information contained in the spreadsheet (see Partial matching rules to learn more about matching partial names).
- grouping A second column must specify which data elements should be analyzed together in a given batch unit: this would be the ID of a single individual when comparing different tissues from the same individual (one individual per batch); or a family name when identifying variants existing within one family (one family per batch).
- Type The third column must specify the type for each data element: the values in this column distinguish tissue samples from controls, or inform about the disease status of a family member (affected/non-affected/proband) when identifying disease causing variants.
Ready-to-use workflows with more than one input in the Biomedical Genomics Workbench fall within two categories; 1) the Somatic Cancer workflow that compares tumor and normal samples, and 2) the Hereditary Disease workflows where a trio or a family of four is analyzed in one workflow.
(Figure 7.8) shows an example of the spreadsheet used in the Somatic Cancer workflows.
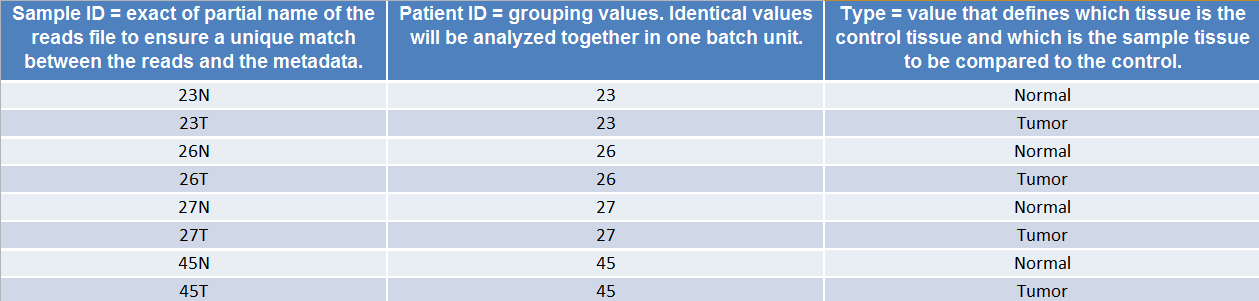

Figure 7.8: Example of a spreadhseet necessary to run a workflow in batch, where the workflow intend to compare two samples.
To launch a workflow with multiple input elements in batch mode:
- Right click on the name of the workflow in the Toolbox panel in the bottom left hand side of the Workbench and select the option "Run in Batch Mode..." (figure 7.9).
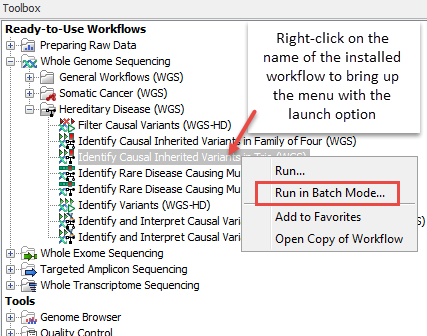
Figure 7.9: The option to "Run in Batch Mode..." appears in the context menu when you right click on the name of an installed workflow that has multiple input elements in the Toolbox panel.A Wizard like that in figure 7.10 should appear.
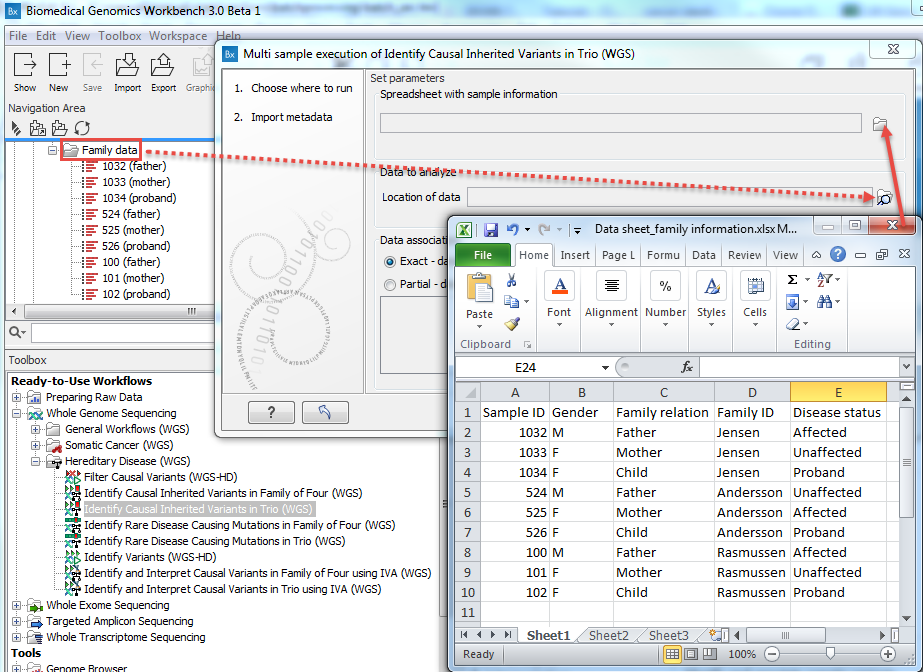
Figure 7.10: Select the information about the data to be analyzed and the folder holding the data to analyze. An example of an Excel sheet with the relevant information is shown. - Select the Excel file containing the information about the data to be analyzed (figure 7.11).
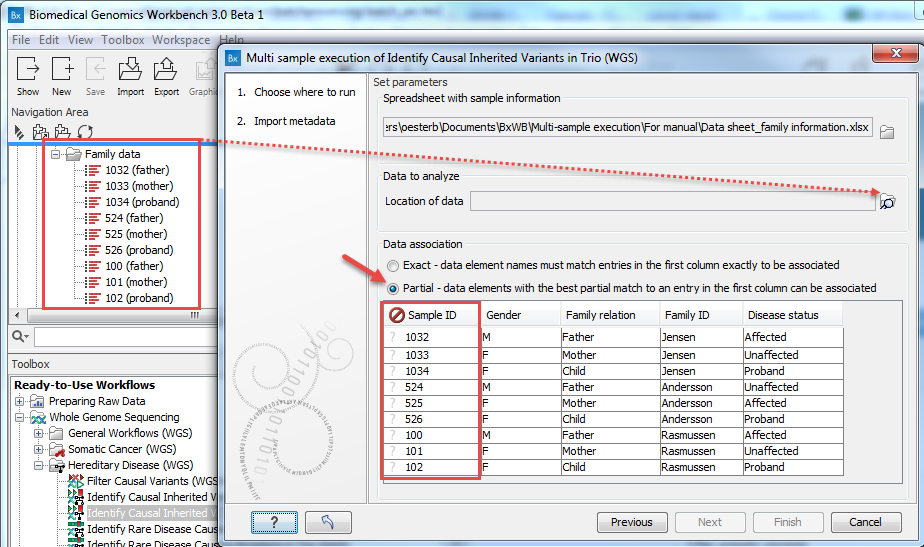
Figure 7.11: When the Excel sheet has been selected, the table found in the lower part of the wizard will show the content of the Excel sheet. The location of the data for this analysis is not yet specified, so a red, no-entry sign is visible in the header of the first column. - Specify the folder with the data, as shown in figure 7.12.
Data elements within the selected folders are considered for the analysis. Subfolders and their contents are not considered unless the subfolder is also selected. Individual data elements cannot be selected.
- Select the appropriate matching scheme - exact or partial. The matching rules applied are the same as those used for metadata association. Exact means that data element names must exactly match an entry in the first column of the Excel file. Partial matching allows for data elements names partially matching an entry in the first column. Partial matching rules are described in detail in section 3.2.2.
An icon with a green check mark (
 ) appears in the table preview next to rows where a data element corresponding to a row of the Excel sheet was uniquely identified. If no match can be made to a given row of the Excel sheet, a question mark (
) appears in the table preview next to rows where a data element corresponding to a row of the Excel sheet was uniquely identified. If no match can be made to a given row of the Excel sheet, a question mark ( ) is displayed.
) is displayed.
Graphical symbols are also presented in the header of the first column of the preview pane to give information about the overall status of the matching of rows in the Excel sheet with data elements in the Workbench:
- When no data elements match information in the Excel sheet, a red, no entry symbol (
 ) is displayed. In this situation, the button labeled Next is not enabled. This is the expected state before any data elements have been selected.
) is displayed. In this situation, the button labeled Next is not enabled. This is the expected state before any data elements have been selected.
- A yellow exclamation mark (
 ) indicates that some, but not all rows in the Excel sheet have been matched to a data element in the selected folder(s).
) indicates that some, but not all rows in the Excel sheet have been matched to a data element in the selected folder(s).
- A green checkmark (
 ) indicates that all rows in the Excel sheet have been matched to a data element in the selected folder(s).
) indicates that all rows in the Excel sheet have been matched to a data element in the selected folder(s).
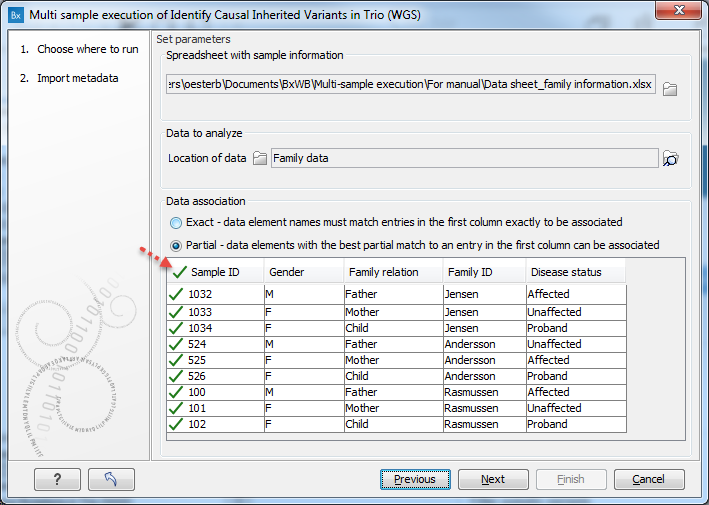
Figure 7.12: Click on a folder or folders that contain the data to be analyzed. Here, the green check mark symbol in the header of the first column in the preview pane indicates that data elements were identified for each of the rows in the Excel sheet. - When no data elements match information in the Excel sheet, a red, no entry symbol (
- Click on the button labeled Next.
In the Grouping area of the Wizard shown in figure 7.13:
- In the Group by drop down menu, select the name of the column containing information that specifies which samples should be analyzed together.
- In the Type drop down menu, select the name of the column containing information that can be mapped to the workflow input type of each data element.
An example is shown in figure 7.13, where a hereditary workflow is being launched in batch mode. Group by is set to a column specifying family names, because each workflow run will analyze a particular family. Type is set to the Disease status column, because the workflow inputs are an unaffected parent, an affected parent and a proband, and the Disease status column holds entries that can be mapped to these input types.
The same Excel sheet shown in figure 7.13 could also be used where the workflow input types were instead mother, father and child. In that case, the column called Family relation would be set as the Type, since that is the column with entries that can be mapped to those particular workflow input types.
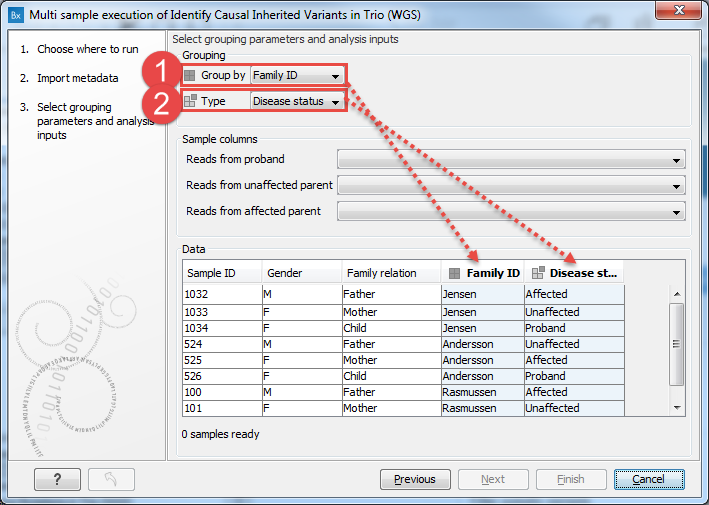
Figure 7.13: A hereditary workflow is being lauched in batch mode. A given workflow run should analyze a family group, so the Group by entry is set to the column Family ID, where family groupings are specified. The workflow input types here are an unaffected parent, an affected parent and a proband. Information that can be mapped to these input types is held in the Disease status column, so this is selected in the Type drop down menu.Further details about the information in the Type column is now entered in the Sample columns area of the Wizard. For each input type for the workflow being launched, a drop down menu is provided containing the column entries from the column specified as containing the Type information.
- For each workflow input type listed, click on the drop down menu and select the term used to identify that particular input type. See figure 7.14.
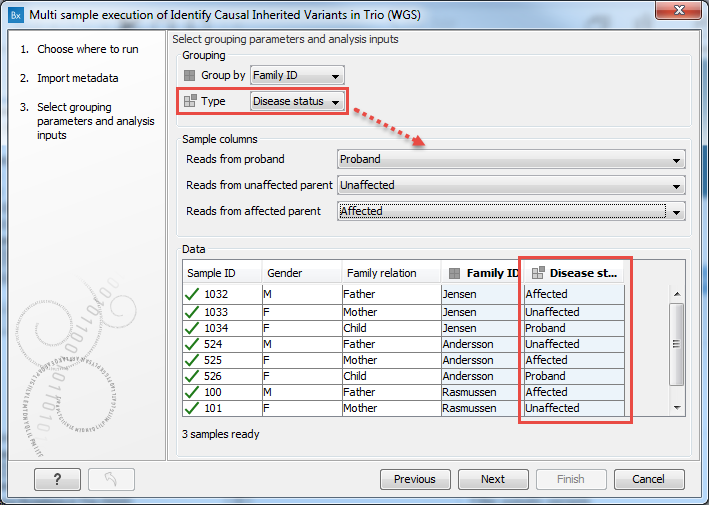
Figure 7.14: The selections shown here indicate that data elements identified as matching rows from the Excel sheet containing "Proband" in the Disease status column should be used as the workflow input type "proband", data elements identified as matching rows containing "Unaffected" should be used as the workflow input type "unaffected parent", and data elements identified as matching rows containing "Affected" should be used as the workflow input type "affected parent". - Click on the button labeled Next.
- Work through any remaining Wizard steps where analysis details are presented and configure any unlocked parameters.
- Choose where to save the outputs of the analysis.
- Click on the button labeled Finish.
Important note: When running the Identify Rare Disease Causing Mutations ready-to-use workflows in batch mode, the gender of all proband samples in a given batch run must be the same. In other words, if multiple families are analyzed in a batch run, the probands must all be female or they must all be male. This is because proband gender is specified as a parameter, and the parameter values provided when setting up a workflow are then used for each analysis in the batch. The same condition applies when running a workflow in batch mode that includes a Trio Analysis. The gender of all child samples being analyzed in a given batch run must be the same.
