The Cell Clonotypes element
A Cell Clonotypes element contains two views, one centered around the identified clonotypes, while the other is centered around the barcodes. Both views contain the following information (see figure 10.2):
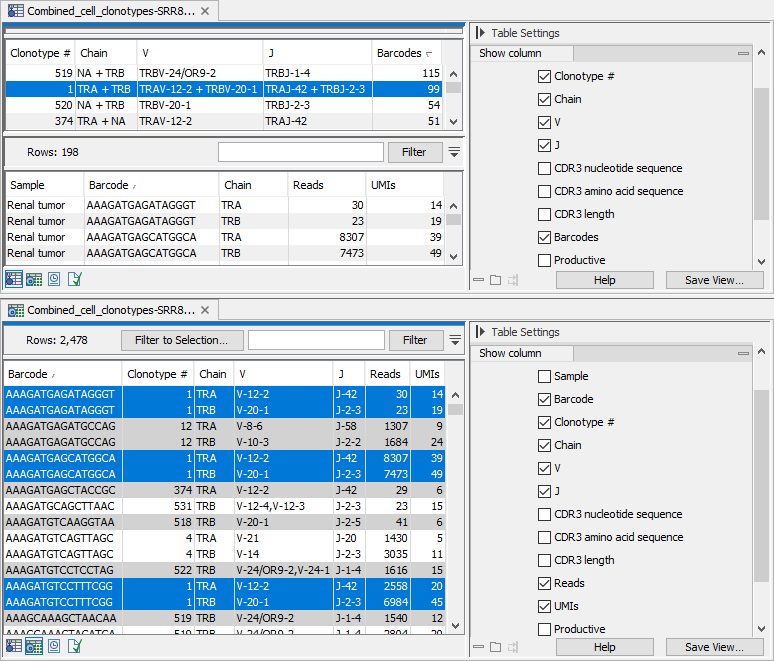
Figure 10.2: Views of the same Cell Clonotypes element. Note that not all table columns are shown. Clonotype with number 1 is highlighted in both views. Top: View centered around the identified clonotypes, sorted after the number of barcodes. The paired clonotypes are shown (both TRA + TRB or TRG + TRD chains), and when just one of the chains could be identified, the missing one is listed as NA, as seen for clonotypes 519 and 374. When a clonotype is selected, a second table lists the barcodes with the corresponding clonotype. Bottom: View centered around the barcodes, sorted by barcode. Rows with the same barcode have the same background color when the table is sorted after the barcode.
- Clonotype #. A unique number for the clonotype.
- Chain: Which of the four gene families
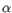 ,
, 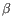 ,
, 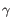 ,
, 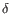 , the clonotype belongs to. To ease filtering, the families are denoted TRA, TRB, TRG and TRD respectively.
, the clonotype belongs to. To ease filtering, the families are denoted TRA, TRB, TRG and TRD respectively.
- V / J. The identified V and J reference segment(s), respectively. If a single unambiguous segment cannot be identified, the segments are separated by a comma.
- CDR3 nucleotide sequence. The nucleotide sequence for CDR3 including the V- and J region-encoded conserved motifs.
- CDR3 amino acid sequence. The translated amino acid sequence for the CDR3 nucleotide sequence, provided that it is in-frame.
- CDR3 length. The length of the CDR3 nucleotide sequence.
- Productive. One of three categories are used to characterize the CDR3 nucleotide sequence:
- Productive. Sequences that are in frame and do not contain a premature stop codon.
- Out-of-frame. Sequences that have a length that is not a multiple of three.
- Premature stop codon. Sequences that contain an in-frame premature stop codon.
Note that the Filter Cell Clonotypes can be used for retaining only the productive clonotypes, see Filter Cell Clonotypes for details.
The view centered around the identified clonotypes additionally contains the numbers of barcodes with the given clonotype. Clicking on a row in this view opens a new table listing the corresponding barcodes (see figure 10.2).
The view centered around the barcodes also provides (see figure 10.2):
- Sample / Barcode. The sample and barcode.
- Reads. The number of reads from the barcode that mapped to the contig from which the specific CDR3 sequence was detected.
- UMIs. The number of unique UMIs the aligned reads correspond to.
