Filter Cell Clonotypes
Sometimes it can be desirable to restrict Cell Clonotypes to only a specific subset, for example only productive clonotypes, or only cells that have both the TRA and TRB chains identified. This can be achieved with the Filter Cell Clonotypes tool, which can be found in the Toolbox here:
Immune Repertoire (![]() ) | Filter Cell Clonotypes (
) | Filter Cell Clonotypes (![]() )
)
The tool takes a Cell Clonotypes (![]() ) element as input and produces a filtered Cell Clonotypes (
) element as input and produces a filtered Cell Clonotypes (![]() ) element.
) element.
The following options can be adjusted (figure 10.3):
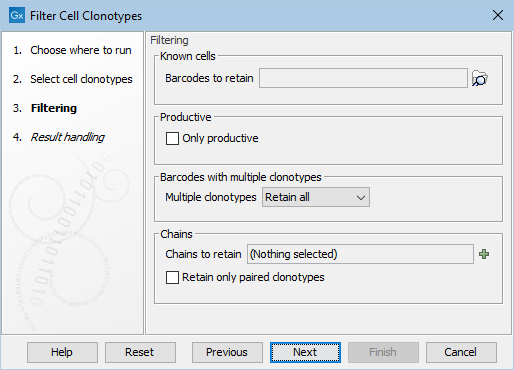
Figure 10.3: The options in the dialog of the Filter Cell Clonotypes tool.
- Barcodes to retain. Multiple elements containing cells can be provided, such as Expression Matrices, Cell Clusters and Cell Annotations. From these, a set of valid cells, identified through the sample and barcode, is obtained as the intersection of the cells in the chosen elements. When used, only the clonotypes for the valid cells are retained in the output.
- Productive. If "Only productive" is enabled, only clonotypes that have a productive CDR3 sequence are retained in the output.
- Barcodes with multiple clonotypes. Barcodes can have more than one clonotype associated with them. Ideally, they contain paired clonotypes for the TRA + TRB chains, or TRG + TRD chains, depending on the cell type. Sometimes a barcode can have fewer or more clonotypes. Configuring the Multiple clonotypes option determines how such barcodes should be handled:
- Retain all. No filter is applied and all clonotypes are retained.
- Retain primary / secondary. When a barcode has more clonotypes for the same chain, only the clonotype with the highest number of reads, or second highest, respectively, is retained. Note that Retain primary also retains all the barcodes containing at most one clonotype per chain, while Retain secondary removes them.
- Retain none. Barcodes containing more than one clonotype for some chain will be removed: only barcodes that have at most one clonotype for each chain are retained.
- Chains to retain. A combination of TRA, TRB, TRG and TRD chains can be chosen and only the clonotypes with the respective chains will be retained. If left empty, no filter is applied.
- Retain only paired clonotypes. Only clonotypes for which a pair is also present are retained. The TRA chain is paired with the TRB chain, and TRG with TRD.
The options above can be mixed and matched to obtain the desired output. Note that the filters are applied in the order given above.
For example, assume we want to only use the primary productive clonotypes with a TRB chain. This can be obtained by enabling "Only productive", setting "Multiple clonotypes" to "Retain primary" and choosing "TRB" in "Chains to retain". If one barcode has two clonotypes with a TRB chain, the primary being non-productive and the secondary being productive, the result will contain the productive chain. Even though this chain was the secondary one in the input, when the tool applies the "Multiple clonotypes" filter, the non-productive clonotype is no longer present in the data and the secondary productive clonotype becomes therefore the primary one.
If a different order for the filters is desired, the tool can be run multiple times with just one of the filter options enabled, in the needed order. For the same example, running the tool first with "Multiple clonotypes" set to "Retain primary" and choosing "TRB" in "Chains to retain", and a second time with "Only productive" enabled, will entirely remove the TRB chains of the mentioned barcode.
The Filter Cell Clonotypes tool can optionally produce a report, summarizing the clonotypes left after filtering. This option is available only when the input element contains just one sample. The output report includes the same information as the report produced by the Single Cell Immune Repertoire Analysis tool, minus the assembly and trimming summaries (see The report output from Single Cell Immune Repertoire Analysis).
