BLAST HSP table
In addition to the graphical display of a BLAST result, it is possible to view the BLAST results in a tabular view. In the tabular view, one can get a quick and fast overview of the results. Here you can also select multiple sequences and download or open all of these in one single step. Moreover, there is a link from each sequence to the sequence at NCBI. These possibilities are either available through a right-click with the mouse or by using the buttons below the table.
The BLAST table view can be shown in the following way:
Click the Show BLAST HSP Table button (![]() ) at the
bottom of the view
) at the
bottom of the view
Figure 26.10 is an example of a BLAST HSP Table.
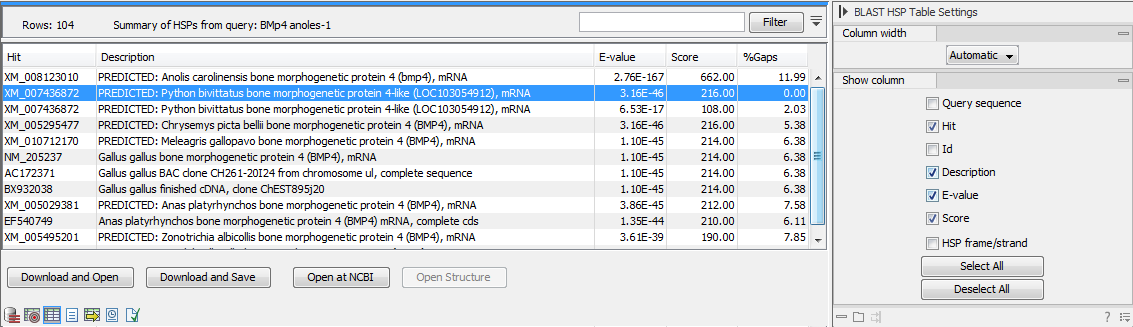
Figure 26.10: BLAST HSP Table. The HSPs can be sorted by the different columns, simply by clicking the column heading.
The BLAST HSP Table includes the following information:
- Query sequence. The sequence which was used for the search.
- HSP. The Name of the sequences found in the BLAST search.
- Id. GenBank ID.
- Description. Text from NCBI describing the sequence.
- E-value. Measure of quality of the match. Higher E-values indicate that BLAST found a less homologous sequence.
- Score. This shows the score of the local alignment generated through the BLAST search.
- Bit score. This shows the bit score of the local alignment generated through the BLAST search. Bit scores are normalized, which means that the bit scores from different alignments can be compared, even if different scoring matrices have been used.
- HSP start. Shows the start position in the HSP sequence.
- HSP end. Shows the end position in the HSP sequence.
- HSP length. The length of the HSP.
- Query start. Shows the start position in the query sequence.
- Query end. Shows the end position in the query sequence.
- Overlap. Display a percentage value for the overlap of the query sequence and HSP sequence. Only the length of the local alignment is taken into account and not the full length query sequence.
- Identity. Shows the number of identical residues in the query and HSP sequence.
- %Identity. Shows the percentage of identical residues in the query and HSP sequence.
- Positive. Shows the number of similar but not necessarily identical residues in the query and HSP sequence.
- %Positive. Shows the percentage of similar but not necessarily identical residues in the query and HSP sequence.
- Gaps. Shows the number of gaps in the query and HSP sequence.
- %Gaps. Shows the percentage of gaps in the query and HSP sequence.
- Query Frame/Strand. Shows the frame or strand of the query sequence.
- HSP Frame/Strand. Shows the frame or strand of the HSP sequence.
In the BLAST table view you can handle the HSP sequences. Select one or more sequences from the table, and apply one of the following functions.
- Download and Open. Download the full sequence from NCBI and opens it. If multiple sequences are selected, they will all open (if the same sequence is listed several times, only one copy of the sequence is downloaded and opened).
- Download and Save. Download the full sequence from NCBI and save it. When you click the button, there will be a save dialog letting you specify a folder to save the sequences. If multiple sequences are selected, they will all open (if the same sequence is listed several times, only one copy of the sequence is downloaded and opened).
- Open at NCBI. Opens the corresponding sequence(s) at GenBank at NCBI. Here is stored additional information regarding the selected sequence(s). The default Internet browser is used for this purpose.
- Open structure. If the HSP sequence contain structure information, the sequence is opened in a text view or a 3D view. Note that the 3D view has special system requirements, see System requirements.
The HSPs can be sorted by the different columns, simply by clicking the column heading. In cases where individual rows have been selected in the table, the selected rows will still be selected after sorting the data.
You can do a text-based search in the information in the BLAST table by using the filter at the upper right part of the view. In this way you can search for e.g. species or other information which is typically included in the "Description" field.
The table is integrated with the graphical view so that selecting a HSP in the table will make a selection on the corresponding sequence in the graphical view.
