View Annotations in sequence views
Figure 13.8
shows an annotation displayed on a sequence.
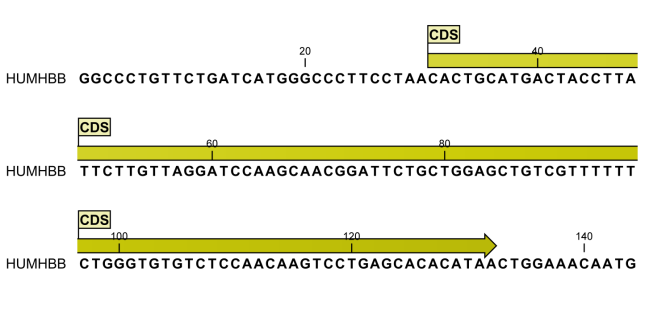
Figure 13.8: An annotation showing a coding region on a genomic dna sequence.
The various sequence views listed in Viewing annotations have different default settings for showing annotations. However, they all have two groups in the Side Panel in common:
- Annotation Layout
- Annotation Types
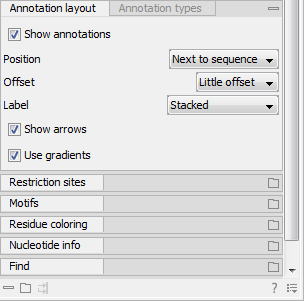
Figure 13.9: The annotation layout in the Side Panel. The annotation types can be shown by clicking on the "Annotation types" tab.
In the Annotation layout group, you can specify how the annotations should be displayed (notice that there are some minor differences between the different sequence views):
- Show annotations. Determines whether the annotations are shown.
- Position.
- On sequence. The annotations are placed on the sequence. The residues are visible through the annotations (if you have zoomed in to 100%).
- Next to sequence. The annotations are placed above the sequence.
- Separate layer. The annotations are placed above the sequence and above restriction sites (only applicable for nucleotide sequences).
- Offset. If several annotations cover the same part of a sequence, they can be spread out.
- Piled. The annotations are piled on top of each other. Only the one at front is visible.
- Little offset. The annotations are piled on top of each other, but they have been offset a little.
- More offset. Same as above, but with more spreading.
- Most offset. The annotations are placed above each other with a little space between. This can take up a lot of space on the screen.
- Label. The name of the annotation can shown as a label. Additional information about the sequence is shown if you place the mouse cursor on the annotation and keep it still.
- No labels. No labels are displayed.
- On annotation. The labels are displayed in the annotation's box.
- Over annotation. The labels are displayed above the annotations.
- Before annotation. The labels are placed just to the left of the annotation.
- Flag. The labels are displayed as flags at the beginning of the annotation.
- Stacked. The labels are offset so that the text of all labels is visible. This means that there is varying distance between each sequence line to make room for the labels.
- Show arrows. Displays the end of the annotation as an arrow. This can be useful to see the orientation of the annotation (for DNA sequences). Annotations on the negative strand will have an arrow pointing to the left.
- Use gradients. Fills the boxes with gradient color.
In the Annotation types group, you can choose which kinds of annotations that should be displayed. This group lists all the types of annotations that are attached to the sequence(s) in the view. For sequences with many annotations, it can be easier to get an overview if you deselect the annotation types that are not relevant.
Unchecking the checkboxes in the Annotation layout will not remove this type of annotations them from the sequence - it will just hide them from the view.
Besides selecting which types of annotations that should be displayed, the Annotation types group is also used to change the color of the annotations on the sequence. Click the colored square next to the relevant annotation type to change the color.
This will display a dialog with five tabs: Swatches, HSB, HSI, RGB, and CMYK. They represent five different ways of specifying colors. Apply your settings and click OK. When you click OK, the color settings cannot be reset. The Reset function only works for changes made before pressing OK.
Furthermore, the Annotation types can be used to easily browse
the annotations by clicking the small button
(![]() ) next to the type. This will display a list
of the annotations of that type (see figure 13.10).
) next to the type. This will display a list
of the annotations of that type (see figure 13.10).
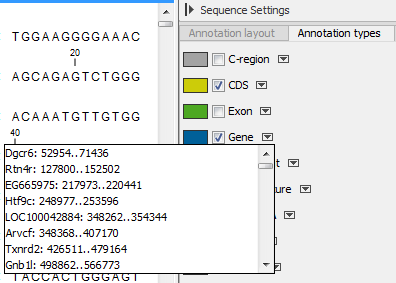
Figure 13.10: Browsing the gene annotations on a sequence.
Clicking an annotation in the list will select this region on the sequence. In this way, you can quickly find a specific annotation on a long sequence.
Note: A waved end on an annotation (figure 13.11) means that the annotation is torn, i.e., it extends beyond the sequence displayed. An annotation can be torn when a new, smaller sequence has been created from a larger sequence. A common example of this situation is when you select a section of a stand-alone sequence and open it in a new view. If there are annotations present within this selected region that extend beyond the selection, then the selected sequence shown in the new view will exhibit these torn annotations.
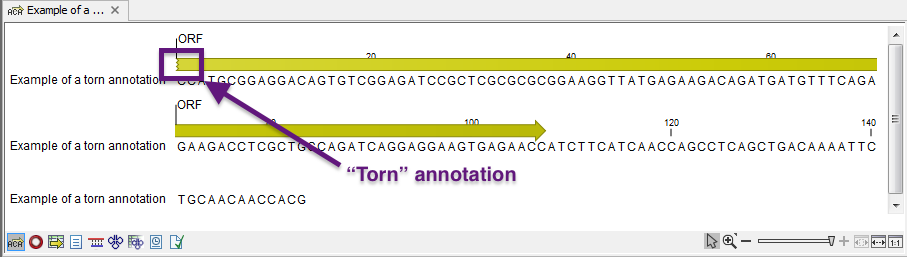
Figure 13.11: Example of a torn annotation on a sequence.
