How does BLAST work?
BLAST identifies homologous sequences using a heuristic method which initially finds short matches between two sequences; thus, the method does not take the entire sequence space into account. After initial match, BLAST attempts to start local alignments from these initial matches. This also means that BLAST does not guarantee the optimal alignment, thus some sequence hits may be missed. In order to find optimal alignments, the Smith-Waterman algorithm should be used (see below). In the following, the BLAST algorithm is described in more detail.
Seeding
When finding a match between a query sequence and a hit sequence, the starting point is the words that the two sequences have in common. A word is simply defined as a number of letters. For blastp the default word size is 3 W=3. If a query sequence has a QWRTG, the searched words are QWR, WRT, RTG. See figure 13.17 for an illustration of words in a protein sequence.
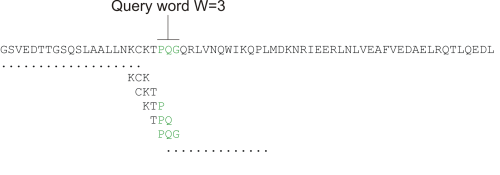
Figure 13.17: Generation of exact BLAST words with a word size of W=3.
During the initial BLAST seeding, the algorithm finds all common words between the query sequence and the hit sequence(s). Only regions with a word hit will be used to build on an alignment.
BLAST will start out by making words for the entire query sequence (see figure 13.17). For each word in the query sequence, a compilation of neighborhood words, which exceed the threshold of T, is also generated.
A neighborhood word is a word obtaining a score of at least T when comparing, using a selected scoring matrix (see figure 13.18). The default scoring matrix for blastp is BLOSUM62. The compilation of exact words and neighborhood words is then used to match against the database sequences.

Figure 13.18: Neighborhood BLAST words based on the BLOSUM62 matrix. Only words where the threshold T exceeds 13 are included in the initial seeding.
After initial finding of words (seeding), the BLAST algorithm will extend the (only 3 residues long) alignment in both directions (see figure 13.19). Each time the alignment is extended, an alignment score is increases/decreased. When the alignment score drops below a predefined threshold, the extension of the alignment stops. This ensures that the alignment is not extended to regions where only very poor alignment between the query and hit sequence is possible. If the obtained alignment receives a score above a certain threshold, it will be included in the final BLAST result.

Figure 13.19: Blast aligning in both directions. The initial word match is marked green.
By tweaking the word size W and the neighborhood word threshold T, it is possible to limit the search space. E.g. by increasing T, the number of neighboring words will drop and thus limit the search space as shown in figure 13.20.
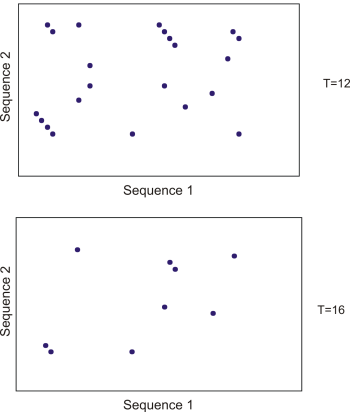
Figure 13.20: Each dot represents a word match. Increasing the threshold of T limits the search space significantly.
This will increase the speed of BLAST significantly but may result in loss of sensitivity. Increasing the word size W will also increase the speed but again with a loss of sensitivity.
