Filter against Known Variants
The Filter against Known Variants tool filters experimental variants based on a known variant track to remove common variants.
Any variant track can be used as the "known variants" track. It may either be produced by the CLC Genomics Workbench, imported or downloaded from variant database resources like dbSNP, 1000 genomes, HapMap etc. (see Import tracks from file and Download Genomes).
To get started, go to:
Toolbox | Resequencing Analysis (![]() ) | Variant Filtering (
) | Variant Filtering (![]() ) | Filter against Known Variants (
) | Filter against Known Variants (![]() )
)
This opens a dialog where you can select a variant track (![]() ) with experimental data that should be filtered.
) with experimental data that should be filtered.
Clicking Next will display the dialog shown in figure 29.1
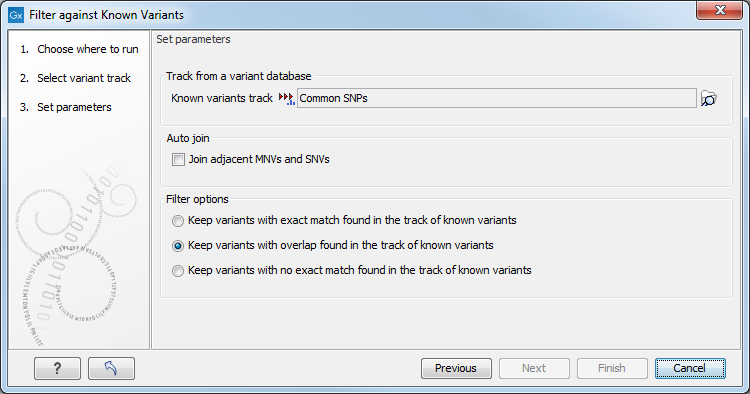
Figure 29.1: Specifying a variant track to filter against.
Select (![]() ) one or more tracks of known variants to compare against. The tool will then compare
each of the variants provided in the input track with the variants in the track of known variants. The output will be a variant track where the remaining variants will depend on the mode of filtering chosen:
) one or more tracks of known variants to compare against. The tool will then compare
each of the variants provided in the input track with the variants in the track of known variants. The output will be a variant track where the remaining variants will depend on the mode of filtering chosen:
- Keep variants with exact match found in the track of known variants. This will filter away all variants that are not found in the track of known variants. This mode can be useful for filtering against tracks with known disease-causing mutations, where the result will only include the variants that match the known mutations. The criteria for matching are simple: the variant position and allele both have to be identical in the input and the known variants track (however, note the extra option for joining adjacent SNVs and MNVs described below). For each variant found, the result track will include information from the known variant. Please note that the exact match criterion can be too stringent, since the database variants need to be reported in the exact same way as in the sample. Some databases report adjacent indels and SNVs separately, even if they would be called as one replacement using the variant detection of CLC Genomics Workbench. In this case, we recommend using the overlap option instead and manually interpret the variants found.
- Keep variants with overlap found in the track of known variants. The first mode is based on exact matching of the variants. This means that if the allele is reported differently in the set of known variants, it will not be identified as a known variant. This is typically not the case with isolated SNVs, but for more complex variants it can be a problem. Instead of requiring a strict match, this mode will keep variants that overlap with a variant in the set of known variants. The result will therefore also include all variants that have an exact match in the track of known variants. This is thus a more conservative approach and will allow you to inspect the annotations on the variants instead of removing them when they do not match. For each variant, the result track will include information about overlapping or strictly matched variants to allow for more detailed exploration.
- Keep variants with no exact match found in the track of known variants. This mode can be used for filtering away common variants if they are not of interest. For example, you can download a variant track from 1000 genomes or dbSNP and use that for filtering away common variants. This mode is based on exact match.
Since many databases do not report a succession of SNVs as one MNV, it is not possible to directly compare variants called with CLC Genomics Workbench with these databases. In order to support filtering against these databases anyway, the option to Join adjacent SNVs and MNVs can be enabled. This means that an MNV in the experimental data will get an exact match, if a set of SNVs and MNVs in the database can be combined to provide the same allele.
Note! This assumes that SNVs and MNVs in the track of known variants represent the same allele, although there is no evidence for this in the track of known variants.
This tool will create a new track where common variants have been removed. The annotations that are left are marked in three different ways:
- Exact match
- This means that the variant position and allele both have to be identical in the input and the known variants track (however, note the extra option for joining adjacent SNVs and MNVs described below).
- Partial MNV match
- This applies to MNVs which can be annotated with partial matches if an SNV or a shorter MNV in the database has an allele sequence that is contained in the allele sequence of the annotated MNV.
- Overlap
- This will report if the known variant track has an overlapping variant.
