The Upload to QCI Interpret tool
The Upload to QCI Interpret tool can be found here:
Tools | QIAseq Panel Expert Tools | QCI Interpretation Integration | Upload to QCI Interpret
Double-click on the Upload to QCI Interpret tool to send a VCF file and possibly metadata information to QCI Interpret.
Select the variant track you want to send (as VCF) to QCI Interpret (figure 8.3), and then select the reference sequence (hg19 by default, but hg38 is also supported). You can then choose which application you would like for the QCI Interpret results (Somatic or Hereditary). Note that the somatic and germline/hereditary workflows of QCI Interpret focus on different end user needs, with somatic being primarily focussed on therapeutic, prognostic, and diagnostic actionability, while germline/hereditary primarily better suited for disease diagnosis/risk.
In case you are working with RNAscan panels analysis, select a Fusions track to upload to QCI Interpret.
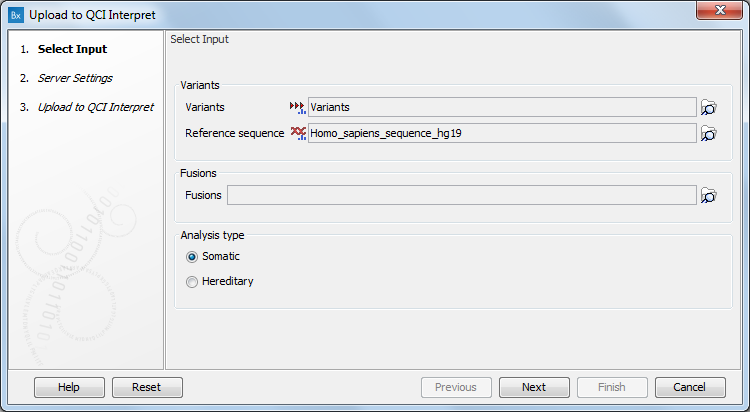
Figure 8.3: The Upload to QCI Interpret dialog.
QCI Interpret configuration requires the user to fill the QCI Interpret dialog (figure 8.4) with the following information:
- Server location. QCI Interpret is hosted in multiple locations. Your QCI Interpret account is created for a specific QCI Interpret server that should be specified here.
- API key ID and API key secret. If you do not know your API credentials, contact support-license@qiagen.com.
- Username. Insert the username associated with the QCI Interpret account.
For additional information on QCI Interpret user account and the QCI Interpret uploader, please refer to the QCI Interpret documentation.
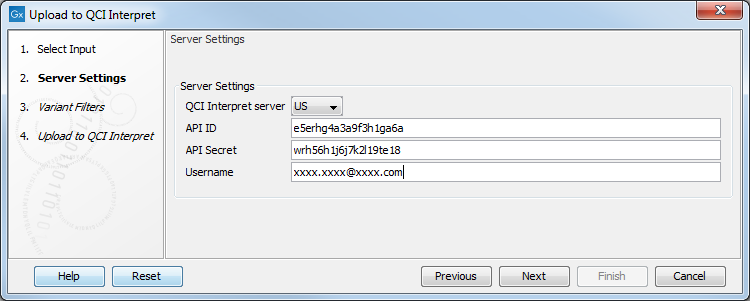
Figure 8.4: Configure the connection to the QCI Interpret server dialog.
In the case of uploading a variant file to QCI Interpret, a dialog called "Variants Filters" will appear next (figure 8.5).
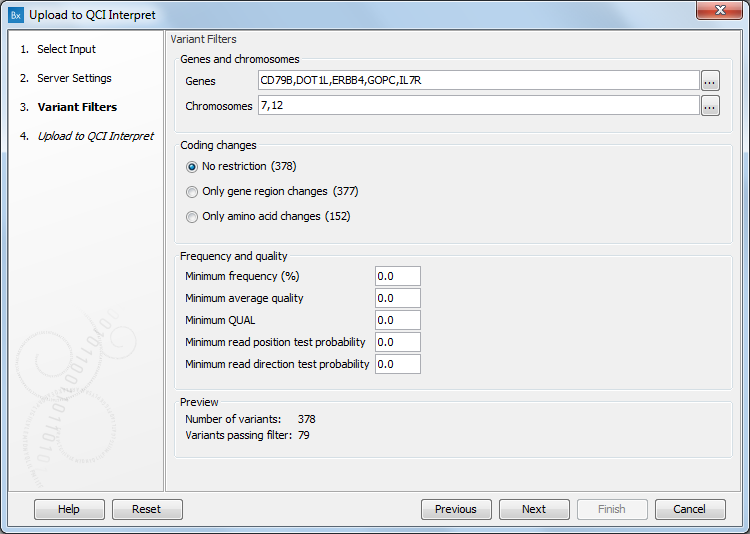
Figure 8.5: The Variants Filters dialog, with the pop up window that allows users to select variants based on their chromosomal location opened as example.
Larger VCF files may need to be filtered to remove the excess variants found during second analysis. Use the next dialog to reduce the number of variants uploaded to QCI Interpret, in particular for somatic pipeline where the upload of variants is currently limited to 400.
Different types of filter are available:
- Genes and chromosomes. Select from the list that opens when clicking to the right of the relevant field all genes or chromosomes of interest. By clicking OK, the pop up window closes and reveals how many variants passed the filtered in the Preview section of the dialog.
- Coding changes. Choose between No restriction, Only gene region changes, Only amino acid changes.
- Frequency and quality. Modify the different filters to reduce the list of variants passing filters (see the Preview section of the dialog).
In the last wizard window, click on Finish to initiate the transfer of variants or fusions to QCI Interpret. The tool will open a browser window with the QCI Interpret interview page. This page must be filled in and the button Submit must be pressed to complete the upload of the files and metadata to QCI Interpret. In cases where a browser window fails to open automatically, the tool will also output a report containing the link to the QCI Interpret interview page.
