Import Known Fusion Information Track
The Import Known Fusion Information Track tool can be found in the Import menu.
The tool takes a TAB separated text file that describes the known fusion information as input (figure 4.18). You can then choose the appropriate reference gene track, saved in the CLC_References folder of the Navigation Area when downloading the QIAseq RNAscan Panels hg38 Reference Data Set.
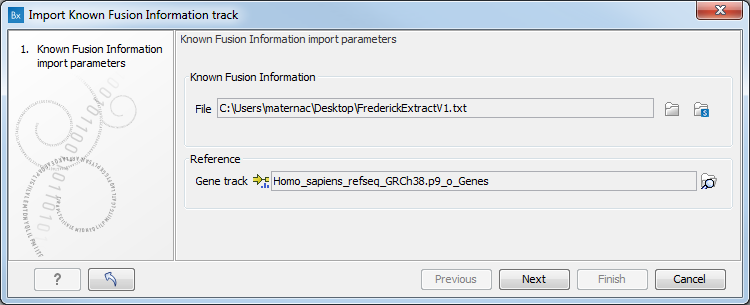
Figure 4.18: Select a text file containing the previously known fusion information, and a reference gene track.
The TAB separated text file format is as follow: The first row must contain the header which defines column names and number of columns in the table. Each subsequent row describes a known fusion information record consisting of a 5'prime and 3'prime gene name, an optional breakpoint position and a list of custom annotations.
The header must include include two specific columns: 5'prime and 3'prime gene names, and two optional columns: 5'prime and 3'prime breakpoint position. Column names are:
- "5prime_gene" : 5'prime gene name.
- "5prime_bp" : 5'prime breakpoint position (optional).
- "3prime_gene" : 3'prime gene name.
- "3prime_bp" : 3'prime breakpoint position (optional).
Any additional columns are added as annotations for the known fusion information record. The order of the columns are not important since the header column names are used to identify the contents of the fusion information.
The 5'prime and 3'prime genes are looked up in the provided Gene track and must therefore be present.
Choose to save in the Navigation Area the newly created track element annotated with (KFI) and recapitulating all the information from the input file. This track can now be used with the Annotate Fusions with Known Fusion Information.
