Compare variants in DNA and RNA
Integrated analysis of genomic and transcriptomic sequencing data is a powerful tool that can help increase our current understanding of human genomic variants. The Compare variants in DNA and RNA ready-to-use workflow identifies variants in DNA and RNA and studies the relationship between the identified genomic and transcriptomic variants.
To run the ready-to-use workflow:
Toolbox | Ready-to-Use Workflows | Whole Transcriptome Sequencing (![]() ) | (Human (
) | (Human (![]() ), Mouse (
), Mouse (![]() ) or Rat (
) or Rat (![]() )) | Compare variants in DNA and RNA (
)) | Compare variants in DNA and RNA (![]() )
)
- Double-click on the Compare variants in DNA and RNA ready-to-use workflow to start the analysis. If you are connected to a server, you will first be asked where you would like to run the analysis. Click on the button labeled Next.
- Select the DNA reads that you would like to analyze (figure 16.8). To select the DNA reads, double-click on the reads file name or click once on the file and then on the arrow pointing to the right side in the middle of the wizard. Click on the button labeled Next.
Figure 16.8: Select the DNA reads to analyze. - Select now the RNA reads to analyze (see figure 16.9).
Figure 16.9: Select the RNA reads to analyze. - Specify a target region for the analysis of the RNA sample with the Indels and Structural Variants tool (figure 16.10).
The targeted region file is a file that specifies which regions have been sequenced. This file is something that you must provide yourself, as this file depends on the technology used for sequencing. You can obtain the targeted regions file from the vendor of your targeted sequencing reagents.
Figure 16.10: Specify the target region for the Indels and Structural Variants tool. - Set the parameters for the Low Frequency Variant Detection step for your RNA sample (see figure 16.11). For a description of the different parameters that can be adjusted in the variant detection step, see http://clcsupport.com/biomedicalgenomicsworkbench/current/index.php?manual=Low_Frequency_Variant_Detection.html.
If you click on "Locked Settings", you will be able to see all parameters used for variant detection in the ready-to-use workflow.
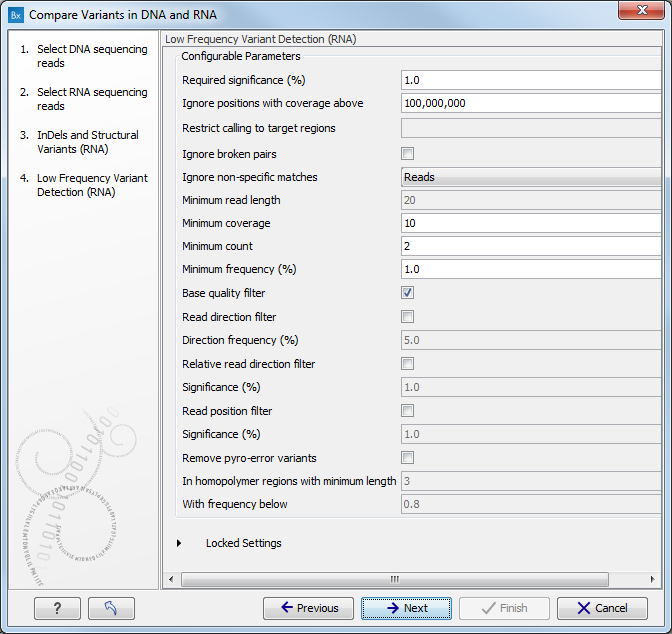
Figure 16.11: Specify the parametes for transcriptomic variant detection. - If you are working with the workflow from the Human folder, specify here the relevant 1000 Genomes population for your RNA sample from the drop-down list (see figure 16.12). Choose the population that matches best the population your samples are derived from.
Under "Locked settings" you can see that "Automatically join adjacent MNVs and SNVs" has been selected. The reason for this is that many databases do not report a succession of SNVs as one MNV as is the case for the Biomedical Genomics Workbench, and as a consequence it is not possible to directly compare variants called with Biomedical Genomics Workbench with these databases. In order to support filtering against these databases anyway, the option to Automatically join adjacent MNVs and SNVs is enabled. This means that an MNV in the experimental data will get an exact match, if a set of SNVs and MNVs in the database can be combined to provide the same allele.
Note! This assumes that SNVs and MNVs in the track of known variants represent the same allele, although there is no evidence for this in the track of known variants.
Figure 16.12: Select the relevant population from the drop-down list. - Repeat the 2 previous steps (or 3 if you are working with the workflow from the human folder) to specify the target region, set the parameters for the Low Frequency Variant Detection the DNA sample - and potentially the population from the 1000 Genomes Project - that characterizes best your DNA sample.
- Click on the button labeled Next to go to the result handling step (figure 16.13).
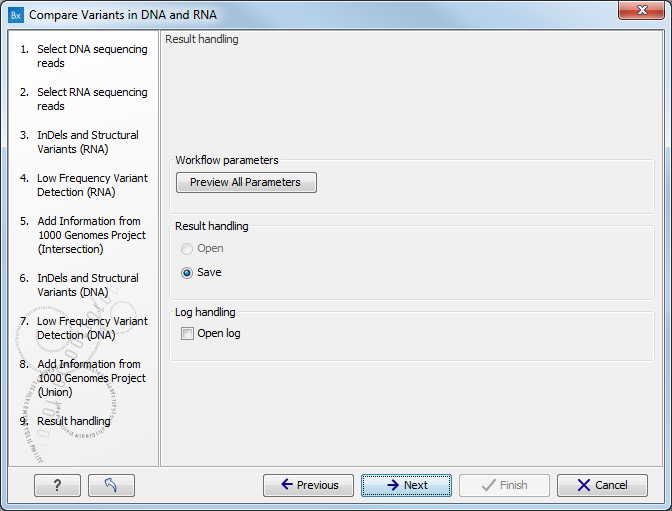
Figure 16.13: Select the relevant population from the drop-down list.Pressing the button Preview All Parameters allows you to preview all parameters. At this step you can only view the parameters, it is not possible to make any changes (see figure 16.14). Choose to save the results and click on the button labeled Finish.
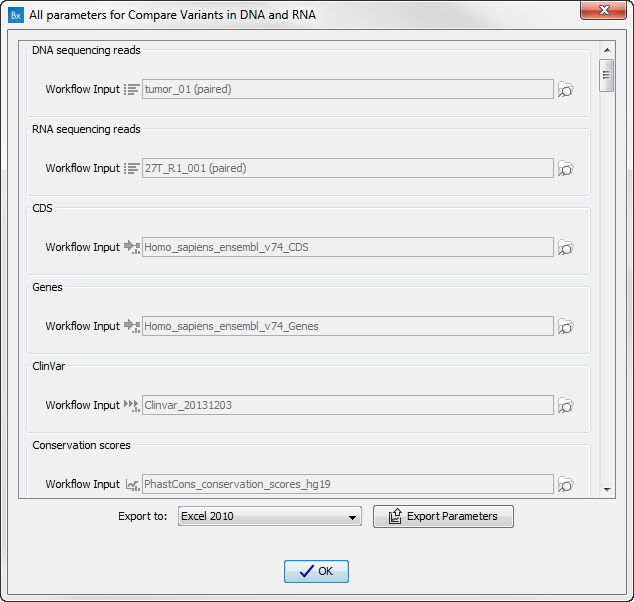
Figure 16.14: Preview all parameters. At this step it is not possible to introduce any changes, it is only possible to view the settings. - Press OK, specify where to save the results, and then click on the button labeled Finish to run the analysis.
Nine different output are generated:
- A DNA Read Mapping and a RNA Read Mapping (
 ) The mapped DNA or RNA sequencing reads. The sequencing reads are shown in different colors depending on their orientation, whether they are single reads or paired reads, and whether they map unambiguously. For the color codes please see the description
in (see http://www.clcsupport.com/biomedicalgenomicsworkbench/current/index.php?manual=View_settings_in_Side_Panel.html).
) The mapped DNA or RNA sequencing reads. The sequencing reads are shown in different colors depending on their orientation, whether they are single reads or paired reads, and whether they map unambiguously. For the color codes please see the description
in (see http://www.clcsupport.com/biomedicalgenomicsworkbench/current/index.php?manual=View_settings_in_Side_Panel.html).
- A DNA Mapping Report and a RNA Mapping Report (
 ) This report contains information about the reads, reference, transcripts, and statistics. This is explained in more detail in the Biomedical Genomics Workbench reference manual in section RNA-Seq report (http://clcsupport.com/biomedicalgenomicsworkbench/current/index.php?manual=RNA_Seq_report.html).
) This report contains information about the reads, reference, transcripts, and statistics. This is explained in more detail in the Biomedical Genomics Workbench reference manual in section RNA-Seq report (http://clcsupport.com/biomedicalgenomicsworkbench/current/index.php?manual=RNA_Seq_report.html).
- An RNA Gene Expression (
 ) A track showing gene expression annotations. Hold the mouse over or right-clicking on the track. If you have zoomed in to nucleotide level, a tooltip will appear with information about e.g. gene name and expression values.
) A track showing gene expression annotations. Hold the mouse over or right-clicking on the track. If you have zoomed in to nucleotide level, a tooltip will appear with information about e.g. gene name and expression values.
- An RNA Transcript Expression (
 ) A track showing transcript expression annotations. Hold the mouse over or right-clicking on the track. A tooltip will appear with information about e.g. gene name and expression values.
) A track showing transcript expression annotations. Hold the mouse over or right-clicking on the track. A tooltip will appear with information about e.g. gene name and expression values.
- A Filtered Variant Track with All Variants Found in DNA or RNA (
 ) This track shows all variants that have been detected in either RNA, DNA or both.
) This track shows all variants that have been detected in either RNA, DNA or both.
- A Filtered Variant Track with Variants Found in Both DNA and RNA (
 ) This track shows only the variants that are present in both DNA and RNA. With the table icon (
) This track shows only the variants that are present in both DNA and RNA. With the table icon ( ) found in the lower left part of the View Area it is possible to switch to table view. The table view provides details about the variants such as type, zygosity, and information from a range of different databases.
) found in the lower left part of the View Area it is possible to switch to table view. The table view provides details about the variants such as type, zygosity, and information from a range of different databases.
- A Genome Browser View Variants Found in DNA and RNA (
 ) A collection of tracks presented together. Shows the annotated variants track together with the human reference sequence, genes, transcripts, coding regions, and variants detected in ClinVar and dbSNP (see figure 16.15).
) A collection of tracks presented together. Shows the annotated variants track together with the human reference sequence, genes, transcripts, coding regions, and variants detected in ClinVar and dbSNP (see figure 16.15).
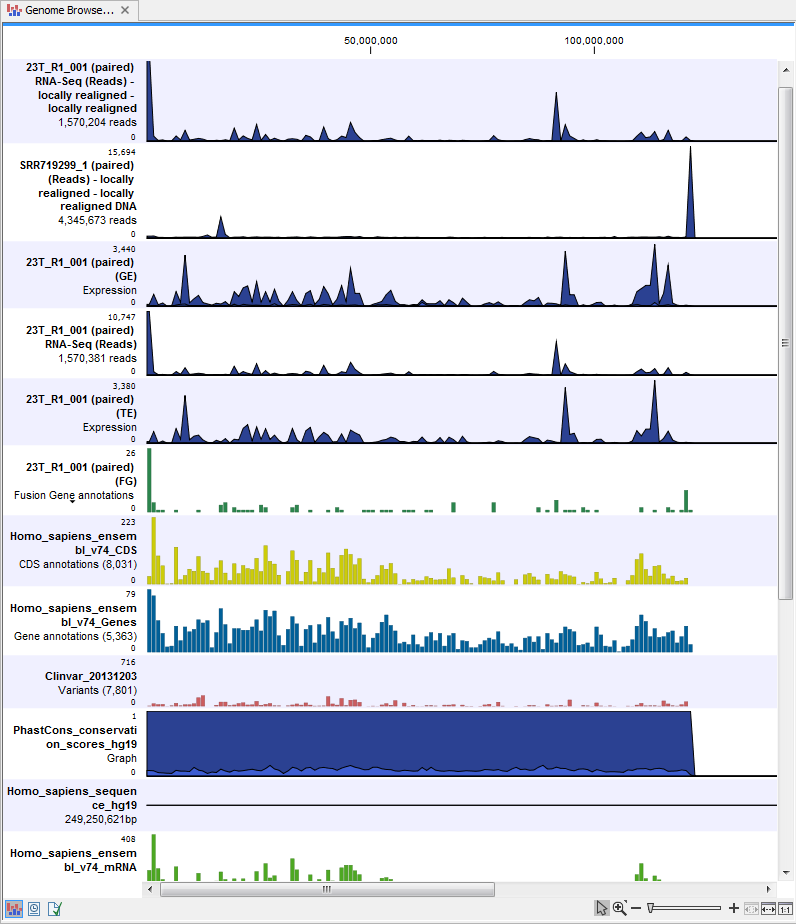
Figure 16.15: The genome browser view makes it easy to compare a range of different data.
The three most important tracks generated are the Variants found in both DNA and RNA track, All variants found in DNA or RNA track, and the Genome Browser View. The Genome Browser View makes it easy to get an overview in the context of a reference sequence, and compare variant and expression tracks with information from different databases. The two other tracks (Variants found in both DNA and RNA track and All variants found in DNA or RNA track) provides detailed information about the detected variants when opened in table view.
