Restriction enzyme lists
Biomedical Genomics Workbench includes all the
restriction enzymes available in the REBASE database, with methylation shown as performed by the cognate methylase rather than by Dam/Dcm. If you want to customize the enzyme database for your installation, see Restriction enzymes database configuration.
However, when performing restriction site analyses, it is often an
advantage to use a customized list of enzymes. In this case, the
user can create special lists containing for example all enzymes available
in the laboratory freezer, or all enzymes used to create a given
restriction map or all enzymes that are available form the preferred
vendor.
In the Example data under Nucleotide->Restriction analysis, there are two enzyme lists: one with the 50 most popular enzymes, and another with all enzymes that are included in the Biomedical Genomics Workbench.
Create enzyme list
Biomedical Genomics Workbench uses enzymes from the REBASE restriction enzyme database at http://rebase.neb.com. If you want to customize the enzyme database for your installation, see Restriction enzymes database configuration.
To create an enzyme list of a subset of these enzymes:
File | New |
Enzyme list (![]() )
)
This opens the dialog shown in figure 32.15
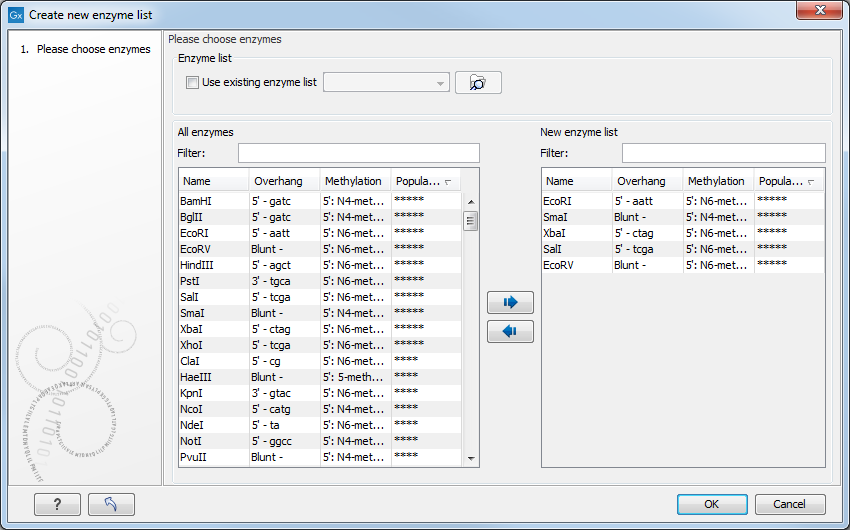
Figure 32.15: Choosing enzymes for the new enzyme list.
Choose which enzyme you want to include in the new enzyme list (see Manage enzymes), and click Finish to open the enzyme list.
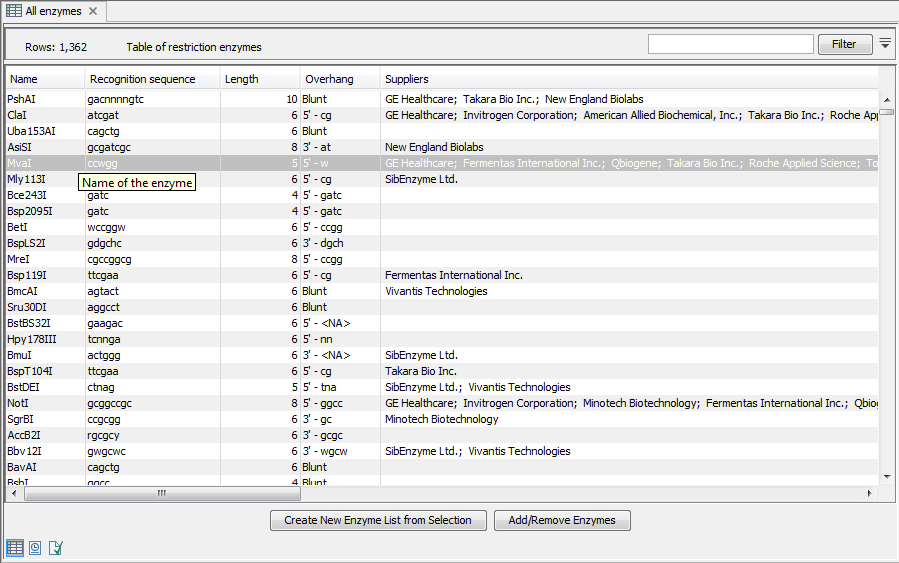
View and modify enzyme list
An enzyme list is shown in figure 32.16. It can be sorted by clicking the columns, and you can use the filter at the top right corner to search for specific enzymes, recognition sequences etc.
If you wish to remove or add enzymes, click the Add/Remove Enzymes button at the bottom of the view. This will present the same dialog as shown in figure 32.15 with the enzyme list shown to the right.
If you wish to extract a subset of an enzyme list, open the list, select the relevant enzymes, right-click on the selection and choose to Create New Enzyme List from
Selection (![]() ).
).
If you combined this method with the filter located at the top of the view, you can extract a very specific set of enzymes. for example, if you wish to create a list of enzymes sold by a particular distributor, type the name of the distributor into the filter and select and create a new enzyme list from the selection.