Remove information from variants
When you use information from various databases to annotate your variants, you may end up with many duplicated annotations or even annotations that you are not really interested in. This tool can help you remove annotations that have been added to variants, so that the results track/results table only includes the information that is of relevance to you.
To run the Remove information from variants, go to the toolbox:
Toolbox | Identify Candidate Variants (![]() ) | Remove information from variants (
) | Remove information from variants (![]() )
)
The input for the tool is an annotated variant track (please make sure that you select a variant track that contains annotations e.g. Amino Acid Change, Exact Match, Conservation Score etc.). If you click on the button labeled "Load Annotations" in the wizard shown in figure 27.2 , the annotations that have been added to the variants in the input track are preloaded in the window below. You can choose, which annotations should be kept or removed. Please use the Ctrl or Shift keys on your keyboard to select the annotations.
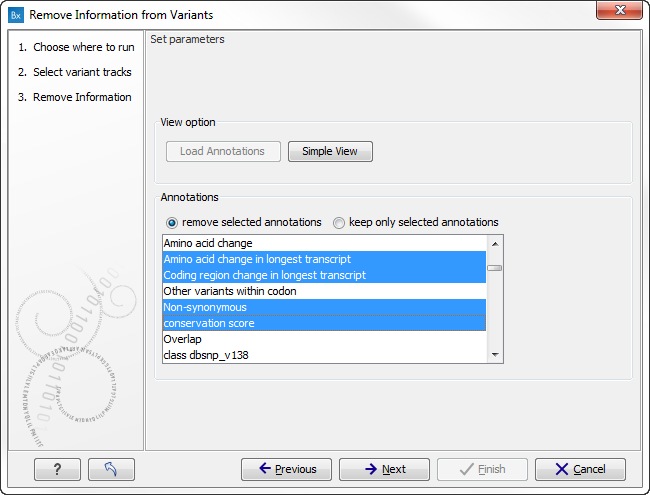
Figure 27.2: The selected annotation columns preload from the
input variant track will be removed in the output variant track.
However, when the tool Remove information from variants is used in a workflow, the option "Load Annotations" is not available and you will have to enter manually the annotation names you want to remove (or that you want to keep, depending on the choice made below in the "to keep or remove" drop-down menu).
To know (and potentially copy) the names of the annotations you need to enter in the window of the workflow wizard, you can start the Remove information from variants tool on its own using as input a similar variant track than the one you will generate with the workflow. Select in the "Load Annotations" menu the annotation names you are interested in removing/keeping. Once you have completed your selection you can click on the button "Simple View" where only the selected annotations will be shown. You can now copy these annotation names, cancel the wizard of the stand alone tool and start the workflow wizard. Paste your selection in the relevant window after having started the workflow wizard.
