How to run the Find Sequence tool
To launch the Find Sequence tool, go to:
Tools | Genome Finishing Module (![]() ) |
Find Sequence (
) |
Find Sequence (![]() )
)
This opens the dialog shown in figure 7.1.
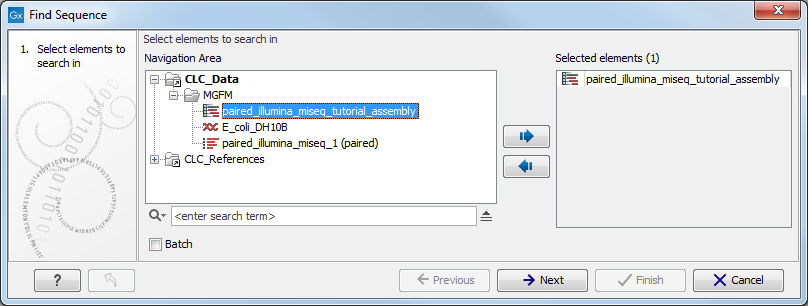
Figure 7.1: Select the elements to search in.
Select the relevant assembled reads and click Next. This leads to the Set search string step shown in figure 7.2.
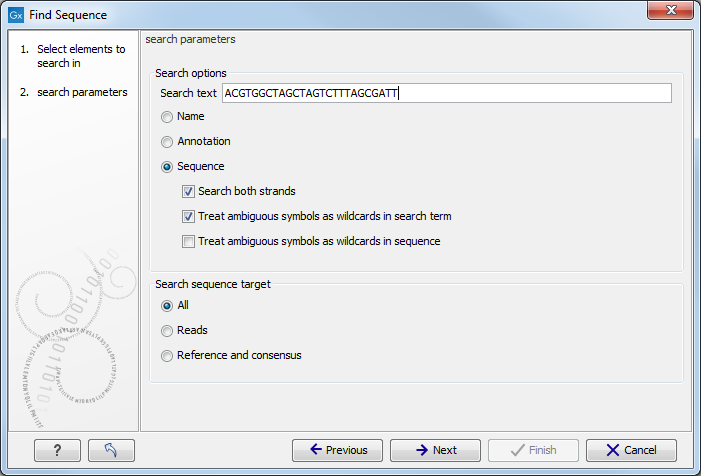
Figure 7.2: Select the parameters for name, sequence or annotation search.
The parameters to be specified in this step are:
- Search text Type or paste the relevant sequence/name that should be used in the search and select whether the search should be performed in a name, sequence or annotation:
- Name. Search for the specified text string in sequence (object) names.
- Annotation. Search for the specified text string in annotations on selected sequences.
- Sequence. Search for the specified text string in selected sequences. When a search is to be performed in a sequence, three new options become available. Tick off the relevant parameters:
- Search both strands
- Treat ambiguous symbols as wildcards in search term
- Treat ambiguous symbols as wildcards in sequence
- Sequence selection
- All sequences. Search for the specified text string in all sequences.
- Reads. Search for the specified text string in only the reads of selected contigs.
- References and consensus. Search for the specified text string in reference and consensus sequence of the selected contigs.
Subsections
