Selecting sequences for
evaluation
Start the Evaluate Structure Hypothesis tool by going to:
Tools | RNA Structure (![]() )|
Evaluate Structure Hypothesis (
)|
Evaluate Structure Hypothesis (![]() )
)
This opens the dialog shown in figure 24.22.
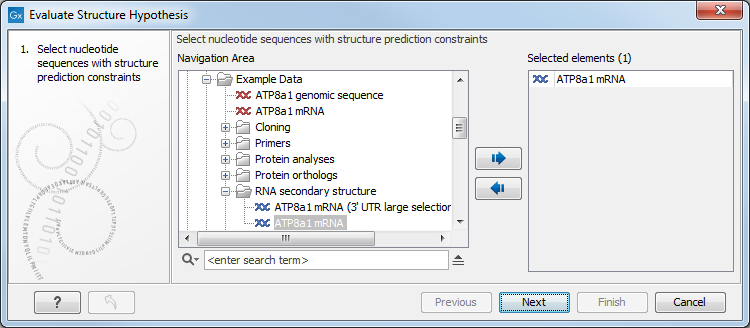
Figure 24.22: Selecting RNA or DNA sequences for evaluating structure hypothesis.
If you had selected sequences before running the tool, those sequences will be listed in the Selected Elements pane of the dialog. Use the arrows to add or remove sequences or sequence lists from the selected elements. Note that the selected sequences must contain a structure hypothesis in the form of manually added constraint annotations.
Click Next to adjust evaluation parameters (see figure 24.23).
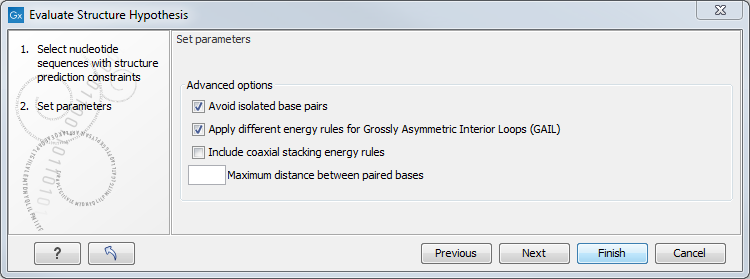
Figure 24.23: Adjusting parameters for hypothesis evaluation.
The partition function algorithm includes a number of advanced options:
- Avoid isolated base pairs. The algorithm filters out isolated base pairs (i.e. stems of length 1).
- Apply different energy rules for Grossly Asymmetric Interior Loops (GAIL). Compute the minimum free energy applying different rules for Grossly Asymmetry Interior Loops (GAIL).
A Grossly Asymmetry Interior Loop (GAIL) is an interior loop that is
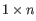 or
or 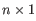 where
where 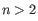 (see
http://mfold.rna.albany.edu/doc/mfold-manual/node5.php)
(see
http://mfold.rna.albany.edu/doc/mfold-manual/node5.php)
- Include coaxial stacking energy rules. Include free energy increments of coaxial stacking for adjacent helices [Mathews et al., 2004].
