Selection of specific nodes
Selection of nodes in a tree is automatically synchronized to the metadata table and the other way around. Nodes in a tree can be selected in three ways:
- Selection of a single node Click once on a single node. Additional nodes can be added by holding down Ctrl (or (
 ) for Mac) and clicking on them (see figure 21.33).
) for Mac) and clicking on them (see figure 21.33).
- Selecting all nodes in a subtree Double clicking on a inner node results in the selection of all nodes in the subtree rooted at the node.
- Selection via the Metadata table Select one or more entries in the table. The corresponding nodes will now be selected in the tree.
It is possible to extract a subset of the underlying sequence data directly through either the tree viewer or the metadata table as follows. Select one or more nodes in the tree where at least one node has a sequence attached. Right click one of the selected nodes and choose Extract Sequence List. This will generate a new sequence list containing all sequences attached to the selected nodes. The same functionality is available in the metadata table where sequences can be extracted from selected rows using the right click menu. Please note that all extracted sequences are copies and any changes to these sequences will not be reflected in the tree.
When analyzing a phylogenetic tree it is often convenient to have a multiple alignment of sequences from e.g. a specific clade in the tree. A quick way to generate such an alignment is to first select one or more nodes in the tree (or the corresponding entries in the metadata table) and then select Align Sequences in the right click menu. This will extract the sequences corresponding to the selected elements and use a copy of them as input to the multiple alignment tool (see section 20.6). Next, change relevant option in the multiple alignment wizard that pops up and click Finish. The multiple alignment will now be generated.
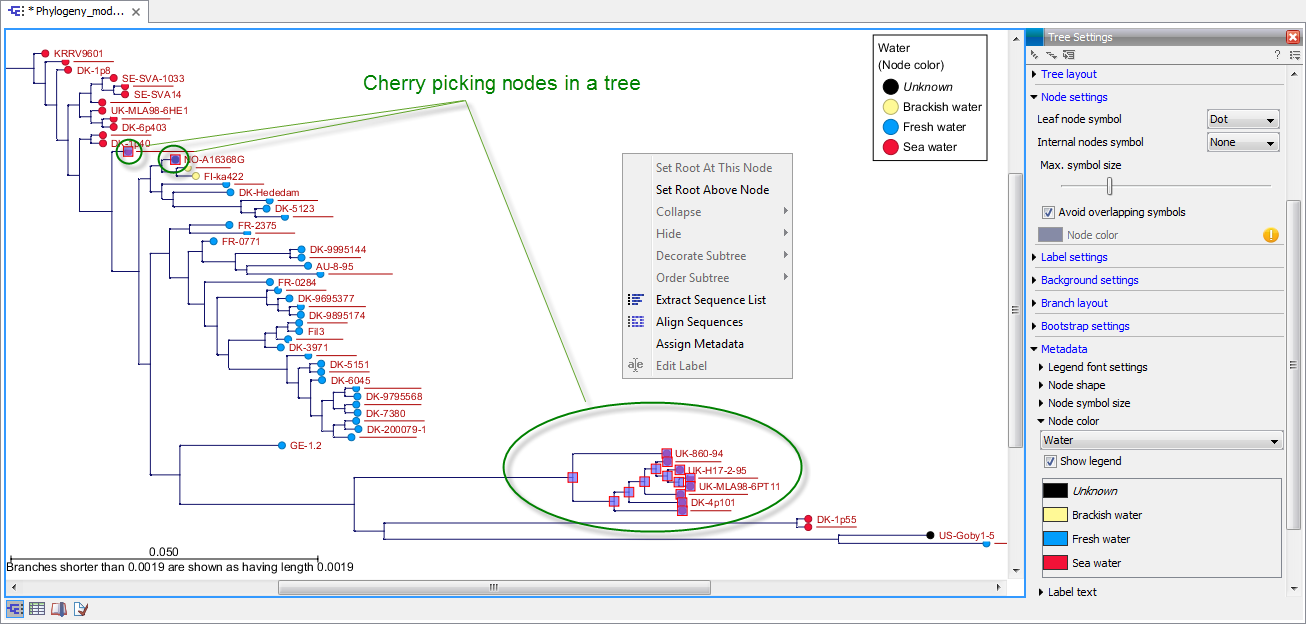
Figure 21.33: Cherry picking nodes in a tree. The selected leaf sequences can be extracted by right clicking on one of the selected nodes and selecting "Extract Sequence List". It is also possible to Align Sequences directly by right clicking on the nodes or leaves.
