Extract Annotations
The Extract annotations tool makes it very easy to extract parts of a sequence (or several sequences) based on its annotations. Using a few steps it is possible to:
- extract e.g. all tRNA genes from the E. coli genome.
- automatically add flanking regions to the annotated sequences.
- search for specific words in all available annotations.
The output is a sequence list that contains sequences carrying the annotation specified (including the flanking regions, if this option was selected).
To extract annotations from a sequence, go to:
Toolbox | Classical Sequence Analysis (![]() ) | General Sequence Analysis (
) | General Sequence Analysis (![]() )| Extract Annotations (
)| Extract Annotations (![]() )
)
This opens the dialog shown in figure 14.1 that asks for either an annotated sequence or an annotation or variant track.
Figure 14.1: Select one or more annotated sequence or annotation or variant tracks.
If you selected tracks as input, the next step will ask for a sequence track to use for extracting the annotations.
Click Next. At the top of the dialog shown in figure 14.2 you can specify a sequence track (in case a track was selected as input), or which annotations to use if an annotated sequence was selected as input:
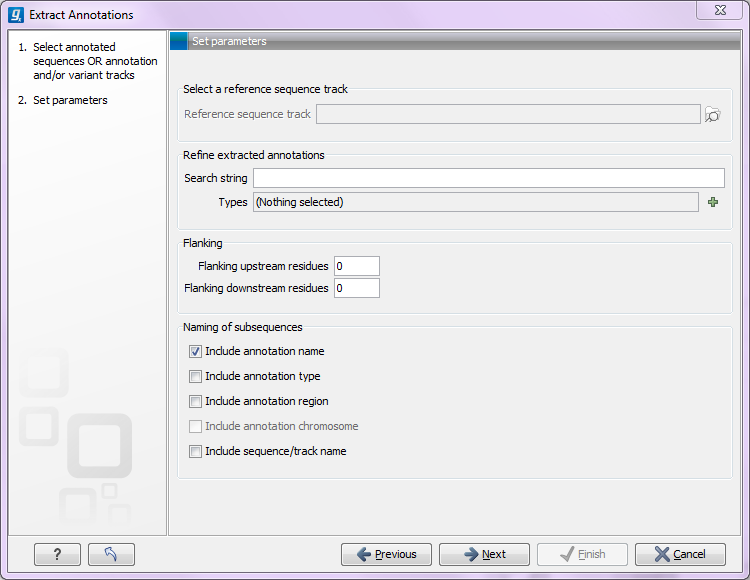
Figure 14.2: Adjusting parameters for extract annotations.
- Search term. All annotations and attached information for each annotation will be searched for the entered term. It can be used to make general searches for search terms such as "Gene" or "Exon", or it can be used to make more specific searches. If you e.g. have a gene annotation called "MLH1" and another called "MLH3", you can extract both annotations by entering "MLH" in the search term field. If you wish to enter more specific search terms, separate them with commas, e.g. "MLH1, Human" will find annotations including both "MLH1" and "Human".
- Annotation types If only certain types of annotations should be extracted, this can be specified here.
The sequence of interest can be extracted with flanking sequences:
- Flanking upstream residues. The output will include this number of extra residues at the 5' end of the annotation.
- Flanking downstream residues. The output will include this number of extra residues at the 3' end of the annotation.
The sequences that are created can be named after the annotation name, type etc:
- Include annotation name. This will use the name of the annotation in the name of the extracted sequence.
- Include annotation type. This corresponds to the type chosen above and will put this information in the name of the resulting sequences. This is useful information if you have chosen to extract "All" types of annotations.
- Include annotation region. The region covered by the annotation on the original sequence (i.e. not including flanking regions) will be included in the name.
- Include sequence/track name. If you have selected more than one sequence as input, this option enables you to discern the origin of the resulting sequences in the list by putting the name of the original sequence into the name of the resulting sequences.
Click Next if you wish to adjust how to handle the results. If not, click Finish.
