Working with isolates
A view of an isolate is shown in figure 4.4.
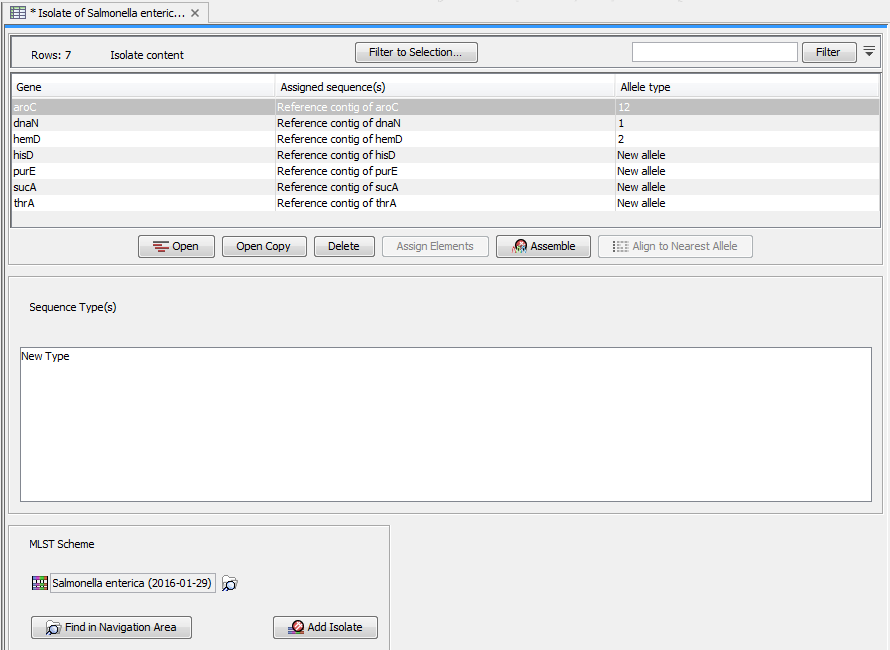
Figure 4.4: An isolate of Streptococcus Uberis typed as number one.
At the top there is a row for each locus. Each row represents a contig which is based on the sequences that have been assigned (either automatically or manually) to this given locus and a reference sequence.
The information in each row is:
- Gene. The name of the locus.
- Assigned sequence(s). The name of the sequences that have been assigned to this locus. In most cases, this will be a reference contig of the selected sequences.
- Allele type. This is the allele type based on comparing the consensus sequence of the contig with the allele sequences in the scheme (this is explained further in Isolate-scheme dependencies).
Subsections
- Isolate-scheme dependencies and the sequence type
- Opening contigs
- Deleting contigs
- Assigning new contigs
- Align to nearest allele
- Assemble to an existing isolate
