Reverse translation parameters
Figure 17.16 shows the choices for making the translation.
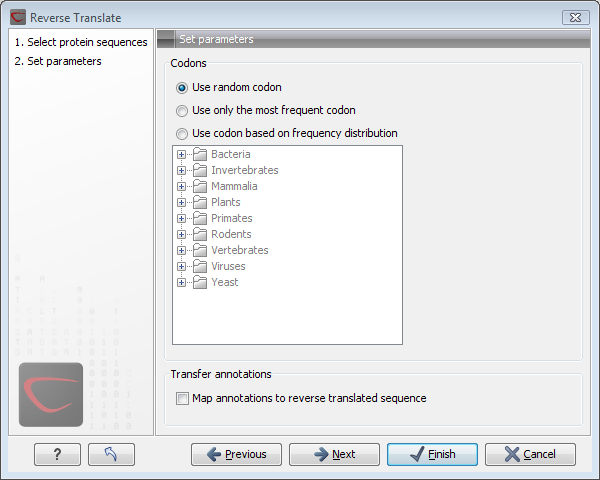
Figure 17.16: Choosing parameters for the reverse translation.
- Use random codon. This will randomly back-translate an amino acid to a codon assuming the genetic code to be 1, but without using the codon frequency tables. Every time you perform the analysis you will get a different result.
- Use only the most frequent codon. On the basis of the selected translation table, this parameter/option will assign the codon that occurs most often. When choosing this option, the results of performing several reverse translations will always be the same, contrary to the other two options.
- Use codon based on frequency distribution. This option is a mix of the other two options. The selected translation table is used to attach weights to each codon based on its frequency. The codons are assigned randomly with a probability given by the weights. A more frequent codon has a higher probability of being selected. Every time you perform the analysis, you will get a different result. This option yields a result that is closer to the translation behavior of the organism (assuming you choose an appropriate codon frequency table).
- Map annotations to reverse translated sequence. If this checkbox is checked, then all annotations on the protein sequence will be mapped to the resulting DNA sequence. In the tooltip on the transferred annotations, there is a note saying that the annotation derives from the original sequence.
The Codon Frequency Table is used to determine the frequencies of the codons. Select a frequency table from the list that fits the organism you are working with. A translation table of an organism is created on the basis of counting all the codons in the coding sequences. Every codon in a Codon Frequency Table has its own count, frequency (per thousand) and fraction which are calculated in accordance with the occurrences of the codon in the organism. The tables provided were made using Codon Usage database http://www.kazusa.or.jp/codon/ that was built on The NCBI-GenBank Flat File Release 160.0 [June 15 2007]. You can customize the list of codon frequency tables for your installation, see Custom codon frequency tables.
Click Next if you wish to adjust how to
handle the results. If not, click Finish.
The newly created nucleotide sequence is shown, and if the analysis was performed on several protein sequences, there will be a corresponding number ofviews of nucleotide sequences. The new sequence is not saved automatically. To save the sequence, drag it into the Navigation Area or press Ctrl + S (![]() + S on Mac) to show the save dialog.
+ S on Mac) to show the save dialog.
