QC for RNAscan Panels
The QC for RNAscan Panels tool can be found in the Toolbox here:
Tools | QIAseq Panel Expert Tools (![]() ) | QIAseq RNAscan Panel Expert Tools (
) | QIAseq RNAscan Panel Expert Tools (![]() ) | QC for RNAscan Panels (
) | QC for RNAscan Panels (![]() )
)
Specify a RNA-seq read mapping as input (figure 6.21).
Figure 6.21: Select a UMI read mapping.
In the next dialog (figure 6.22), specify the mRNA track and the primer track that are saved in the CLC_References folder of the Navigation Area when downloading the QIAseq RNAscan Panels hg38 Reference Data Set. You can also set a maximal distance between a read and a primer start for them to be considered matching. It is set by default to 0, which means that a read will not be considered as starting in the primer unless it maps exactly to the start of the primer.
Figure 6.22: Specify mRNA and primer tracks.
The tool outputs a primer track with annotated read coverage and a report that recapitulates QC data (figure 6.23). The primer track gives information about each primer, as well as their read coverage, whether they overlap with target or housekeeping genes.
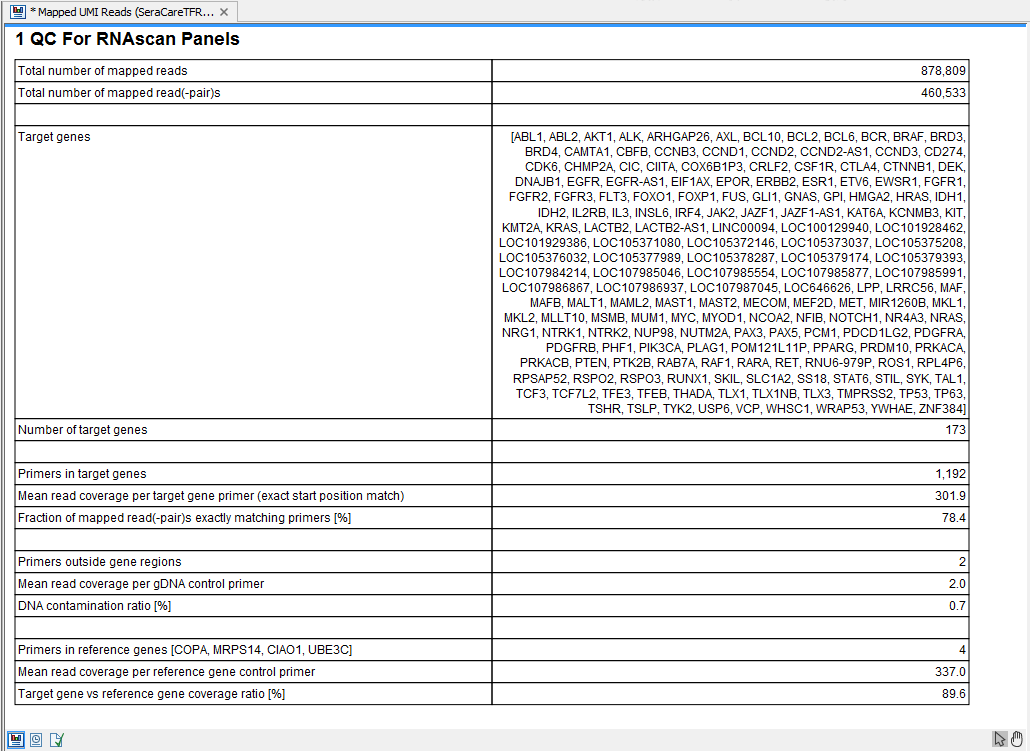
Figure 6.23: QC for RNAscan Panels report.
The QIAseq RNAscan Panels Report
A QC for RNAscan Panels Report contains the following information:
- Total number of mapped reads, counting paired reads as two.
- Total number of mapped reads(-pair)s, counting paired reads as one.
- Target genes: genes targeted by the primers.
- Number of target genes.
- Primers in target genes: how many primers are within the target genes regions.
- Mean read coverage per target gene primer (exact start position match): coverage from reads that start exactly at the primer start site.
- Fraction of mapped read(-pair)s exactly matching primers [%].
- Primers outside gene regions: gDNA control primers outside target gene regions, used for detection of DNA contamination of samples.
- Mean read coverage per non-gene primer: a number higher than 0 indicates DNA contamination of the sample. A mean gDNA coverage around 50 reads or higher may increase false positive signal level.
- DNA contamination [%]: calculated using the Mean read coverage per target gene primer and the Mean read coverage per non-gene primer metrics. In general, if the Mean read coverage per non-gene primer metric is around 50 or higher, the chances for a false positive may be higher.
- Primers in reference genes [COPA, MRPS14, CIAO1, UBE3C]: QIAseq RNAscan panels typically include four reference gene primers.
- Mean read coverage per reference gene control primer: the reference genes should have a mean coverage of at least 300 reads, otherwise the effective input is too low, and false negatives are expected.
- Target gene versus reference gene coverage ratio [%]
