Annotate Variants with Effect Scores (WES)
The Annotate Variants with Effect Scores (WES) ready-to-use workflow takes a variant track as input, and outputs a variant track with effect scores annotations added to non-reference SNVs.
See Annotate with Effect Scores for information about the underlying tool used to add these annotations.
Run the Annotate Variants with Effect Scores (WES) workflow
- The Annotate Variants (WES) workflow can be found in the Toolbox at:
Ready-to-Use Workflows | Whole Exome Sequencing (
 ) | General Workflows (WES) (
) | General Workflows (WES) ( ) | Annotate Variants with Effect Scores (WES) (
) | Annotate Variants with Effect Scores (WES) ( )
)
- In the first wizard step (figure 19.8), select the input variant track.
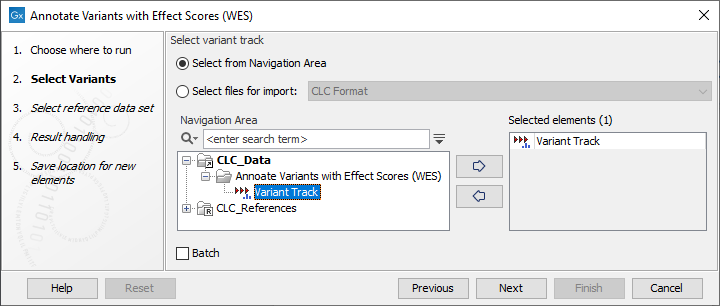
Figure 19.8: Select the relevant variant track to annotate. - In the next wizard step (figure 19.9), select the effect score data.
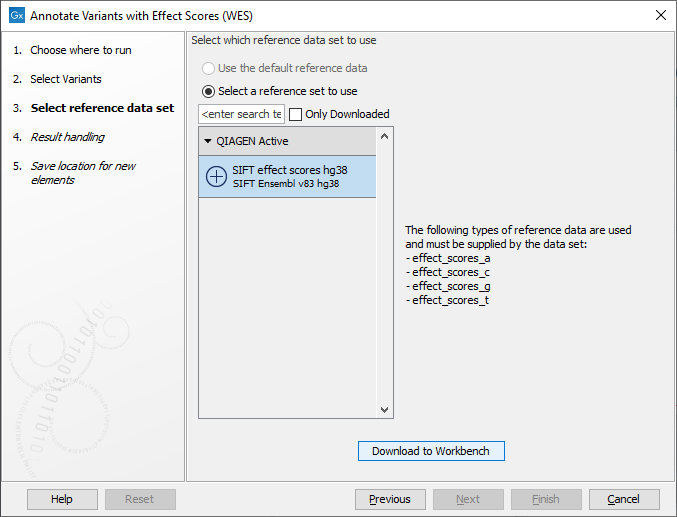
Figure 19.9: Choose the relevant reference data set. - In the last wizard step you can check all the workflow settings by clicking on the button labeled Preview All Parameters.
If you wish to make changes to unlocked parameters, click on the Previous button until you get back to the relevant wizard step.
- Choose to Save your results and click on the button labeled Finish.
Output from the Annotate Variants with Effect Scores (WES) workflow
The Annotate Variants with Effect Scores (WES) outputs a variant track (![]() ) with the original annotations, plus the effect score annotations on non-reference SNVs. Effect scores may not be available for particular positions,
since some effect scores can only be computed for coding regions.
) with the original annotations, plus the effect score annotations on non-reference SNVs. Effect scores may not be available for particular positions,
since some effect scores can only be computed for coding regions.
When hovering the mouse cursor over an individual variant, a tooltip appears containing detailed information about that variant, include the effect score, where available.
The effect score values can also be reviewed by opening the table view of the variant track, as shown in figure 19.10.
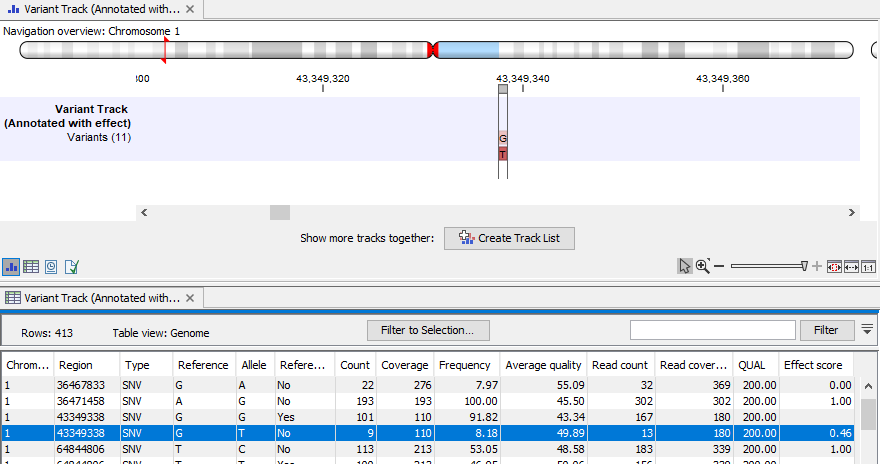
Figure 19.10: Variants annotated with effect scores, here shown in a split view with the track view at the top and the table view at the bottom.
