Annotate with Effect Scores
The Annotate with Effect Scores tool annotates SNV variants with precomputed effect scores. An effect score indicates the impact of a mutation on the gene or transcript. Synonymous mutations have low impact, whereas mutations introducing premature stop-codons or altering protein structures have high impact. Typically, scores are given in a range from 0 to 1, where 1 indicates completely neutral mutations and 0 indicates deleterious mutations, the range and interpretation does however depend on the score used.
To run the Annotate with Effect Scores tool, go to:
Toolbox | Resequencing Analysis (![]() ) | Variant Annotation (
) | Variant Annotation (![]() ) | Annotate with Effect Scores (
) | Annotate with Effect Scores (![]() )
)
Select the variant track (![]() ) as input, when you click Next you will need to provide four tracks with effect scores (see figure 22.2).
The track Effect score A contains the precomputed effect scores of SNVs where the new allele is A.
The other three tracks are defined similarly.
) as input, when you click Next you will need to provide four tracks with effect scores (see figure 22.2).
The track Effect score A contains the precomputed effect scores of SNVs where the new allele is A.
The other three tracks are defined similarly.
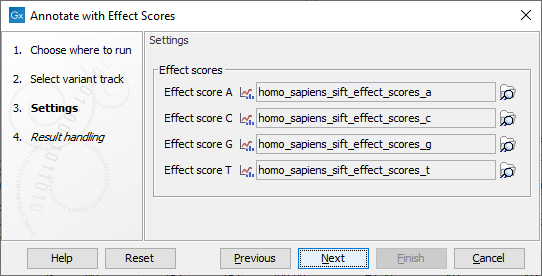
Figure 22.2: Select Effect Score tracks.
The tool outputs a variant track with an Effect score annotation added.
