De Novo Assemble Long Reads and Polish with Short Reads
The De Novo Assemble Long Reads and Polish with Short Reads workflow performs de novo assembly of long reads and polishes the assembly with high-quality short reads. The workflow works best with uncorrected long reads why it is not recommended to run the Correct Long Reads tool before running this workflow.
Launching the workflow
To run the workflow, go to:
Template Workflows (![]() ) | Long Read Workflows (
) | Long Read Workflows (![]() ) | De Novo Assemble Long Reads and Polish with Short Reads (
) | De Novo Assemble Long Reads and Polish with Short Reads (![]() )
)
Launch the workflow and step through the wizard.
- Select the long reads to be assembled (figure 2.1).
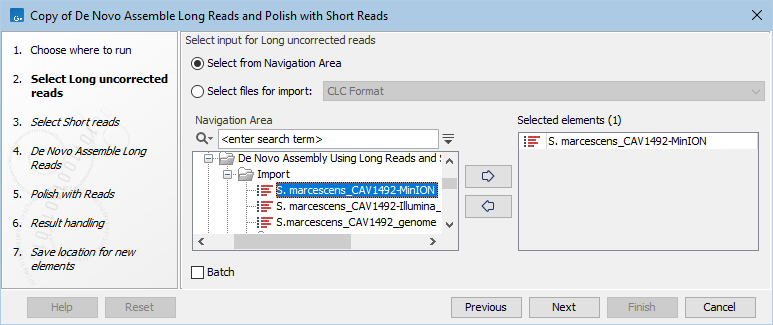
Figure 2.1: Select long, uncorrected reads to assemble - Select the short reads to be used for polishing (figure 2.2).
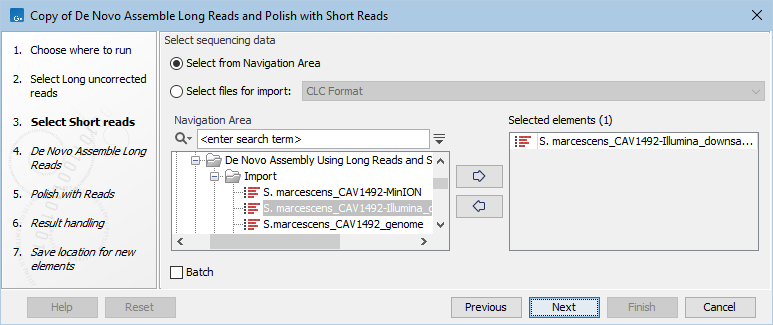
Figure 2.2: Select short reads used to polish the assembly - If you have more than one sample (set of long and short reads) to assemble, check the Batch checkbox in the above steps. For further details on how to run workflows in batch mode when you have more than one input, see http://resources.qiagenbioinformatics.com/manuals/clcgenomicsworkbench/current/index.php?manual=Batching_workflows_with_more_than_one_input_changing_per_run.html.
- Check the PacBio HiFi checkbox if your long reads are PacBio HiFi reads (figure 2.3).
Figure 2.3: Check the checkbox if the long reads are PacBio HiFi reads. - Check the Include unpolished sequences to include unpolished contigs in the output contig list (figure 2.4).
Figure 2.4: Select whether to keep unpolished contigs. - In the final step, choose a location to save the results to.
Workflow tools and outputs
The De Novo Assemble Long Reads and Polish with Short Reads workflow contains five tool (figure 2.5):
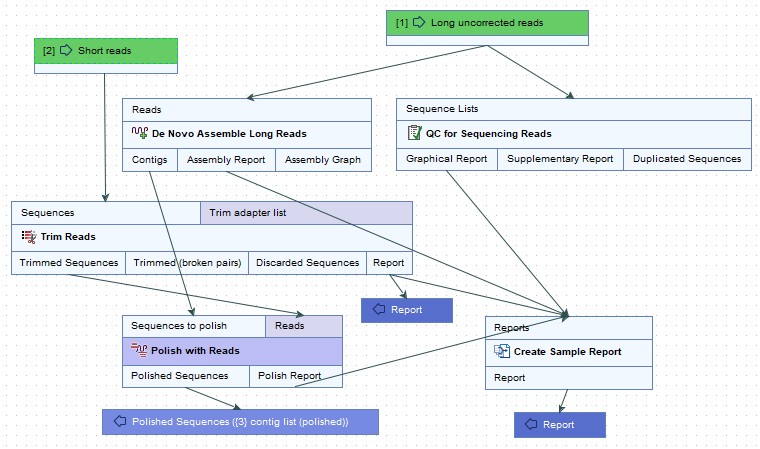
Figure 2.1: The De Novo Assemble Long Reads and Polish with Short Reads workflow.
- QC for Sequencing Reads. See http://resources.qiagenbioinformatics.com/manuals/clcgenomicsworkbench/current/index.php?manual=QC_Sequencing_Reads.html.
- Trim Reads. See http://resources.qiagenbioinformatics.com/manuals/clcgenomicsworkbench/current/index.php?manual=Trim_Reads.html.
- De Novo Assemble Long Reads. See De Novo Assemble Long Reads.
- Polish with Reads. See Polish with Reads.
- Create Sample Report. See http://resources.qiagenbioinformatics.com/manuals/clcgenomicsworkbench/current/index.php?manual=Create_Sample_Report.html.
The outputs provided by the workflow are:
- Graphical QC trim report.
- Contig list (polished). The output from Polish with Reads. Depending on the workflow settings, this may contain unpolished reads.
- Sample report. Contains general information about the sample and contigs produced. Includes QC for raw reads, assembly and polishing statistics.
