Noise filters
These filter thresholds specify when a filter is applied to a genome element for it to be disregarded as noise (figure 5.4):- Minimum coverage: Only variants in regions covered by at least this many reads are called.
- Maximum coverage: All positions with coverage above this value will be ignored when inspecting the read mapping for variants. The option is highly useful in cases where you have a read mapping which has areas of extremely high coverage as are areas around centromeres for example.
- Minimum average base quality: Alleles that have an average base quality below this threshold are disregarded as noise.
- Read direction filter: The read direction filter removes alleles that are almost exclusively present in either forward or reverse reads. For many sequencing protocols such alleles are most likely to be the result of amplification induced errors. Note, however, that the filter is NOT suitable for amplicon data, as for this you will not expect coverage of both forward and reverse reads. The filter has a single parameter:
- Direction frequency: A filter is applied to alleles that are not supported by at least this frequency of reads from each direction.
- Relative read direction filter: The relative read direction filter attempts to do the same thing as the 'Read direction filter', but does this in a statistical, rather than absolute, sense: it tests whether the distribution among forward and reverse reads of the variant carrying reads is different from that of the total set of reads covering the site. The statistical, rather than absolute, approach makes the filter less stringent. The filter has one parameter:
- Significance: A filter is applied to alleles whose read direction distribution is significantly different from the expected with a test at this level. The lower you set the significance cut-off, the fewer alleles will be filtered out.
- Read position filter: The read position filter is a filter that attempts to remove systematic errors in a similar fashion as the 'Read direction filter', but that is also suitable for hybridization-based data. It removes alleles that are located differently in the reads carrying it than would be expected given the general location of the reads covering the variant site. This is done by categorizing each sequenced nucleotide (or gap) according to the mapping direction of the read and also where in the read the nucleotide is found; each read is divided in five parts along its length and the part number of the nucleotide is recorded. This gives a total of ten categories for each sequenced nucleotide and a given site will have a distribution between these ten categories for the reads covering the site. If a distinct allele is present in the site, you would expect the allele nucleotides to follow the same distribution. The read position filter carries out a test for whether the read position distribution of the allele carrying reads is different from that of the total set of reads covering the site. The filter has one parameter:
- Significance: A filter is applied to alleles whose read position distribution is significantly different from the expected with a test at this level. The lower you set the significance cut-off, the fewer alleles will be filtered out.
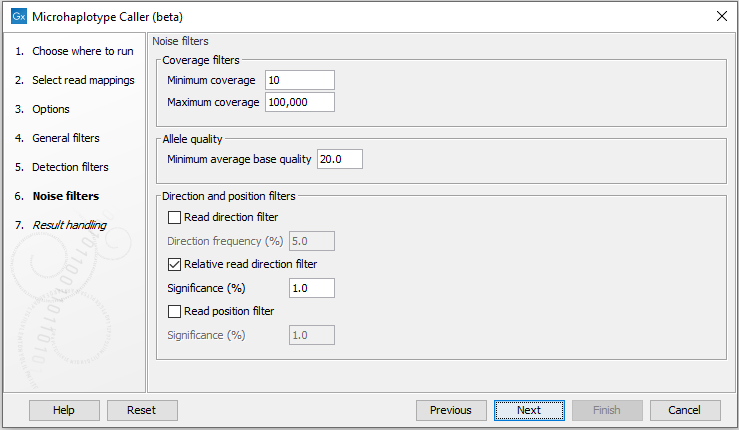
Figure 5.4: The Microhaplotype Caller noise filters.
