Filter Cell Clonotypes
Sometimes it can be desirable to restrict TCR Cell Clonotypes (The Filter Cell Clonotypes tool can be found in the Toolbox here:
Toolbox | Single Cell Analysis (![]() ) | Immune Repertoire (
) | Immune Repertoire (![]() ) | Filter Cell Clonotypes (
) | Filter Cell Clonotypes (![]() )
)
The tool takes a Cell Clonotypes element as input and produces a filtered element.
The following options can be adjusted (figure 13.11):
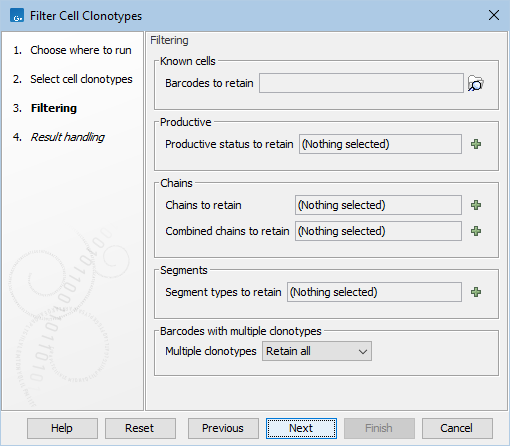
Figure 13.11: The options in the dialog of the Filter Cell Clonotypes tool.
- Barcodes to retain. Multiple elements containing cells can be provided, such as Expression Matrices, Cell Clusters and Cell Annotations. From these, a set of valid cells, identified through the sample and barcode, is obtained as the intersection of the cells in the chosen elements. When used, only the clonotypes for the valid cells are retained in the output.
- Productive status to retain. A mixture of `Productive', `Out of frame' and `Premature stop codon' can be chosen and only the clonotypes with the respective productive status will be retained. If left empty, no filter is applied.
- Chains to retain. A mixture of:
- for TCR Cell Clonotypes: TRA, TRB, TRG and TRD;
- for BCR Cell Clonotypes: IGH, IGK and IGL;
- Combined chains to retain. A mixture of:
- for TCR Cell Clonotypes: TRA + TRB and TRG + TRG;
- for BCR Cell Clonotypes: IGH + IGK and IGH + IGL;
- Segment types to retain. A mixture of `V', `D', `J' and `C' can be chosen and only the clonotypes that have identified segments for all respective segment types will be retained. This means that, for example, if `D' is chosen, only chains for which the D segment is used will be retained, and for those chains, only the clonotypes for which the identification of the D segment was successful will be retained. If left empty, no filter is applied.
- Multiple clonotypes. Barcodes can have more than one clonotype associated with them, see Primary and secondary clonotypes. Different types of filters can be chosen:
- Retain all. No filter is applied and all clonotypes are retained.
- Retain primary. Only the primary clonotypes are retained.
- Retain secondary. Only the secondary clonotypes are retained.
- Retain primary and secondary. Both the primary and the secondary clonotypes are retained.
- Retain none. Only barcodes containing just primary clonotypes are retained.
The options above can be mixed and matched to obtain the desired output. Note that the filters are applied in the order given above.
For example, assume we want to only use the primary TRB clonotypes with D segments. This can be obtained by setting "Chains to retain" to "TRB", "Segment types to retain" to "D", and "Multiple clonotypes" to "Retain primary". If one barcode A has a primary TRB clonotype without D segments and a secondary TRB clonotype with D segments, the former will be removed first and the second with D segments will become the primary one. Hence, "Retain primary" will have no effect on this barcode. If another barcode has two TRB clonotypes with D segments, "Retain primary" will remove the secondary clonotype.
If the desired behavior is that barcode A should be entirely removed from the output, as its primary TRB clonotype does not have D segments, the tool can be run multiple times such that the filters are applied in a different order. By running the tool with "Multiple clonotypes" set to "Retain primary" first, the barcode will have only the clonotype without D segments. A second execution of the tool with "Segment types to retain" set to "D" will entirely remove the barcode.
The Filter Cell Clonotypes tool can optionally produce a report for each sample found in the input element, summarizing the clonotypes left after filtering. The output report includes the same information as the report produced by the Single Cell V(D)J-Seq Analysis tool, minus the assembly and trimming summaries (see The report output from Single Cell V(D)J-Seq Analysis).
To obtain a report summarizing clonotypes across samples, see Compare Single Cell Immune Repertoires.
