Count-based and extra-chromosomal filters
Low quality barcodes can arise for various reasons, such as damaged cells or library preparation problems. Potential low quality barcodes can be identified by detecting outliers in the distributions of the metrics listed below. An outlier is a barcode with a value for a given metric that is more than three median absolute deviations (MADs) from the median value.
- Number of reads. Barcodes with few number of reads result from losing RNA during library preparation.
- Number of expressed features. Barcodes with few expressed features indicate that the diverse transcript population has not been successfully captured.
- Proportion of reads mapped to spike-in control regions. When spike-in controls are used, barcodes with proportionally many reads mapped to the spike-in controls are symptomatic of loss of endogenous RNA, as the same amount of spike-in RNA should have been added to each cell.
- Proportion of reads mapped to features indicative of low quality. Barcodes with proportionally many reads mapped to certain features are indicative of low quality cells. For example, loss of cytoplasmic RNA from perforated cells can lead to high expression of mitochondrial genes in eukaryotes [Islam et al., 2014,Ilicic et al., 2016].
Count-based filters
In this dialog of QC for Single Cell, the filters using the total number of reads and expressed features can be enabled and customized.
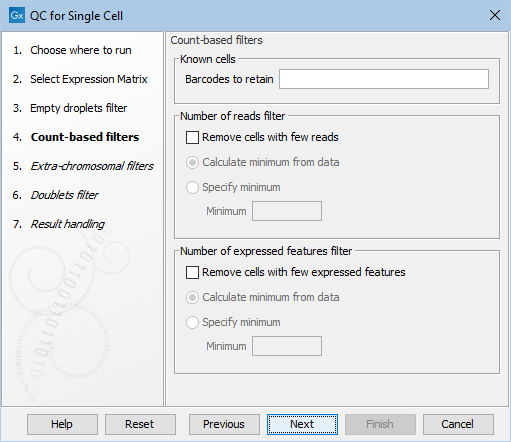
Figure 7.6: The default settings in the Count-based filters dialog.
The dialog first allows for manually specifying a list of barcodes to be retained as cells in Barcodes to retain (figure 7.6). These would typically be barcodes that are otherwise removed by any of the filters applied. See more details in Choosing barcodes to retain.
The following options can be adjusted for the Count-based filters (figure 7.6):
- Remove cells with few reads. Enables filtering based on the total number of reads.
- Remove cells with few expressed features. Enables filtering based on the total number of expressed features.
- For both filters, the outlier detection can be fine-tuned by selecting:
- Calculate minimum from data. Outliers are detected as being more than three MADs below the median.
- Specify minimum. Outliers are detected as being below the threshold specified in the Minimum parameter. This can be useful when the metric distribution is not normal.
Extra-chromosomal filters
In this dialog of QC for Single Cell, the filters using the proportion of reads mapped to spike-in controls and features indicative of low quality reads can be enabled and customized.
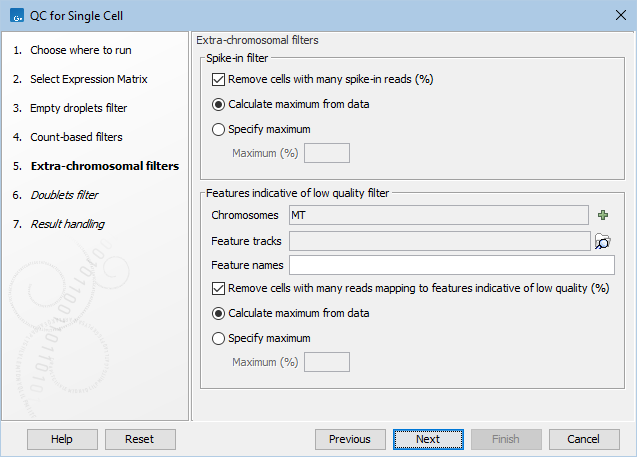
Figure 7.7: The default settings in the Extra-chromosomal filters dialog.
The following options can be adjusted in the Extra-chromosomal filters dialog (figure 7.7):
- Remove cells with many spike-in reads (%). Enables filtering based on the proportion of reads mapped to spike-in controls.
- Chromosomes. The name of the mitochondria chromosome and/or other chromosomes containing only features indicative of low quality cells. Can be left empty or multiple chromosomes can be chosen.
- Feature tracks. Feature tracks containing only features indicative of low quality cells. Can be left empty or multiple tracks can be chosen.
- Feature names. Names or ids features indicative of low quality cells. Any white-space characters, and ",", and ";" are accepted as separators.
- Remove cells with many reads mapping to features indicative of low quality (%). Enables filtering based on the proportion of reads mapped to features indicative of low quality, as defined through Chromosomes, Feature track, and/or Feature names. It requires that at least one of these options is set.
- For both filters, the outlier detection can be fine-tuned by selecting:
- Calculate maximum from data. Outliers are detected as being more than three MADs above the median.
- Specify maximum. Outliers are detected as being above the threshold specified in the Maximum (%) parameter. This can be useful when the metric distribution is not normal.
