Preliminary steps to run the "Type a Known Species" workflow
Before starting the workflow,
- Download the NCBI microbial genomes using the Download Bacterial Genomes from NCBI tool (see the Download Bacterial Genomes from NCBI section). Note that this is a very large file and it takes several hours to download it.
- Download the MLST schemes using the Download MLST Schemes tool (see the Downloading schemes section).
- Download the Database for the Find Resistance tool using the Download of Database for Find Resistance tool (see the Download Database for Find Resistance section).
- Create a New Result Metadata table using the Create Result Metadata Table tool (see the Create Result Metadata Table section).
When you are ready to start the workflows, your navigation area should look similar to the figure 7.2.
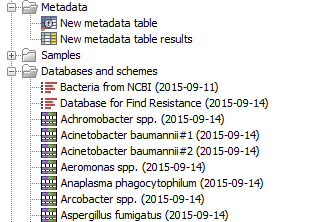
Figure 7.2: Overview of the Navigation area after creating the result metadata table and downloading the databases and MLST schemes necessary to run the workflows.
