Download Bacterial Genomes from NCBI
Download the NCBI collection of bacterial genomes representatives directly from NCBI's FTP site with the Download Bacterial Genomes from NCBI tool. Note! The NCBI download of all bacterial genomes may take at least a few hours depending on your bandwidth. If you are interested in only a few genomes, it is possible to add a filter before starting the tool
To import directly from NCBI's FTP site:
Toolbox | Microbial Genomics Module (![]() ) | Typing and Epidemiology (beta) (
) | Typing and Epidemiology (beta) (![]() ) | Download Bacterial Genomes from NCBI (
) | Download Bacterial Genomes from NCBI (![]() )
)
This will open the following wizard window (figure 9.1):
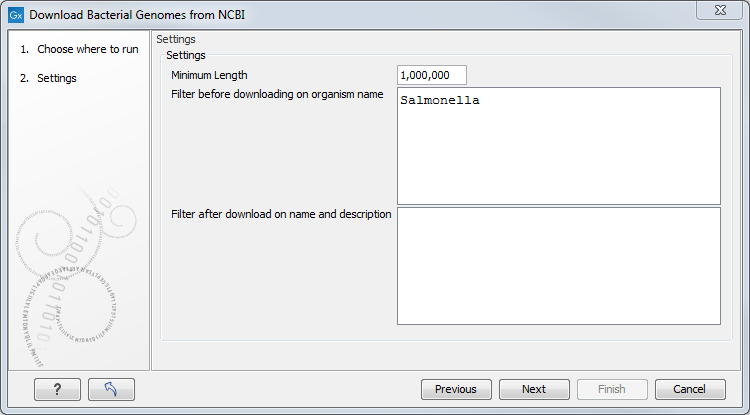
Figure 9.1: An example of downloading only Salmonella from the NCBI's bacterial genomes database.
The settings are:
- Minimum Length: is by default set to 1,000,000 nt to favor bacterial genomes instead of plasmids.
- Filter before downloading on organism name: is by default empty, but by defining one or more taxon, it is possible to limit the download to a subset of the whole NCBI's bacterial genomes database.
- Filter after download on name and description: is by default empty, but by defining one or more phrases, one can limit the output to those sequences which include at least one of these phrases in their description or their name.
Specify a location to save the database. We recommend to use a folder where you save all the databases and MLST schemes necessary to run some of the Microbial Genomics Module tools. Click Finish.
The imported database includes a list of different bacterial genome sequences as well as the associated accession numbers (acc nos), descriptions, taxonomy and size of the sequences.
Note! After download, it is always possible to select a subset of bacterial genomes and saving the reduced list in a separate file (see Extracting a subset of NCBI's bacterial genomes database). This can reduce significantly subsequent analysis runtime.
