Download Pathogen Reference Databases
The Download Pathogen Reference Databases tool is similar to the Download Bacterial Genomes from NCBI, but easier to use in that it will download at once the latest release of the following species-specific pathogen reference genome databases from the NCBI Pathogen Detection Project:
- Salmonella enterica
- Listeria monocytogenes
- Escherichia coli and Shigella
- Campylobacter jejuni
- Acinetobacter baumannii
- Klebsiella pneumoniae
Each reference genome will be annotated with the following metadata (when available):
- serovar
- strain
- taxonomy
- sample collection date
- geographical location
- isolation source
- host
- host disease
- outbreak
- SRA run id
- SRA project id
To run the tool, go to:
Toolbox | Microbial Genomics Module (![]() ) | Typing and Epidemiology (beta) (
) | Typing and Epidemiology (beta) (![]() ) | Download Pathogen Reference Databases (
) | Download Pathogen Reference Databases (![]() )
)
In the first window (figure 9.2), choose the database(s) you want to download.
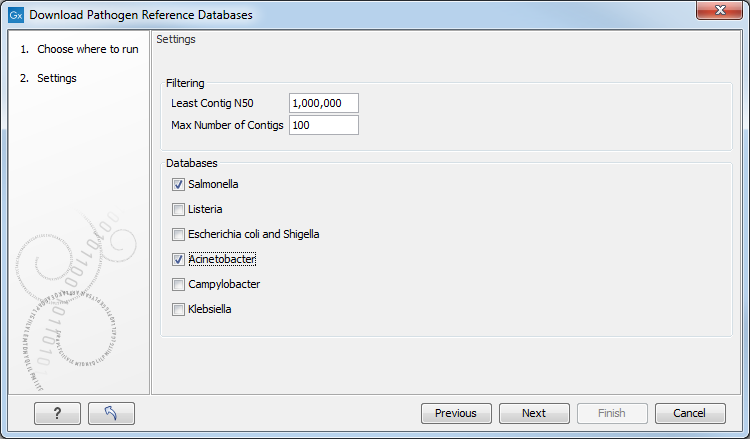
Figure 9.2: Downloading the Actinobacter and Salmonella Pathogen References Databases using the filtering default values.
To ensure high enough quality of the database content, it is possible to filter the pathogen reference genomes being downloaded based on:
- The least amount of contig N50 (the default value is set at 1,000,000 bp)
- The maximum number of contigs (the default value is set at 100)
The time it will take to download the data depends on how many databases are downloaded and the bandwidth of your internet connection.
You can choose to generate a report with the following summary data for each downloaded database:
- Number of reference genomes before filtering
- Number of reference genomes after filtering
- Number of output sequences
- Database version number
- Date of release of the database on the NCBI Pathogen Detection project's ftp site
- Sequence metadata statistics
- Sequence taxonomy statistics, including the amount of entries without taxonomy information.
It is possible to edit the missing or incorrect taxonomic entries in the sequence list table manually by right-clicking on the field to be added or edited, or by using the Set Up Pathogen Reference Database tool (see section). It is also possible to merge multiple reference genome databases to add genomes of interest that are not present in the online database, as well as to delete a reference genome from a pathogen reference genome database when references are not relevant to the analysis or when they are of poor quality. The date of each full database download, as well as the ftp address of the root of the source NCBI Pathogen Detection Project database will be included in the log of the tool and in the history of the downloaded files such that users can locate the downloaded files on NCBI's ftp site again at a later time.
