Insert one sequence into another
Sequences can be inserted into each
other in several ways as described in
Manipulate sequences. When you
chose to insert one sequence into another you will be presented with
a dialog where all sequences in the view are present (see
figure 20.10).
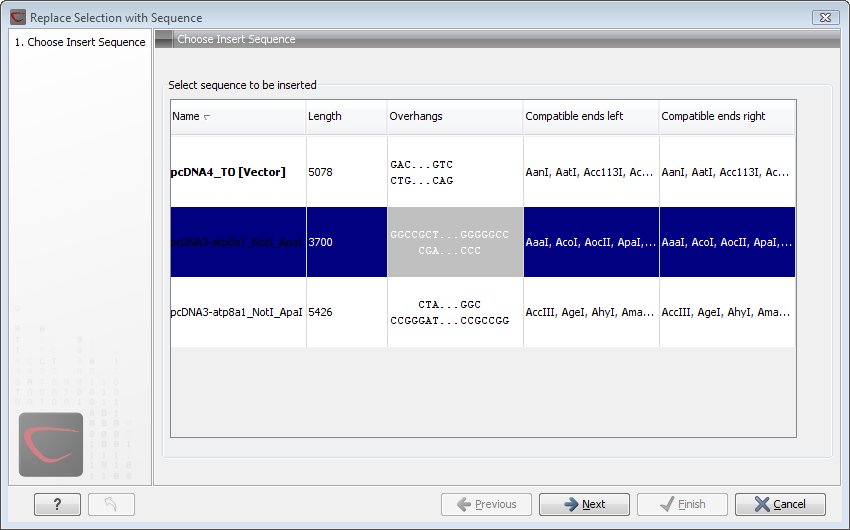
Figure 20.10: Select a sequence for insertion.
The sequence that you have chosen to insert into will be marked with bold and the text [vector] is appended to the sequence name. Note that this is completely unrelated to the vector concept in the cloning workflow.
The list furthermore includes the length of the fragment, an indication of the overhangs, and a list of enzymes that are compatible with this overhang (for the left and right ends, respectively). If not all the enzymes can be shown, place your mouse cursor on the enzymes, and a full list will be shown in the tool tip.
Select the sequence you wish to insert and click Next.
This will show the dialog in figure 20.11).
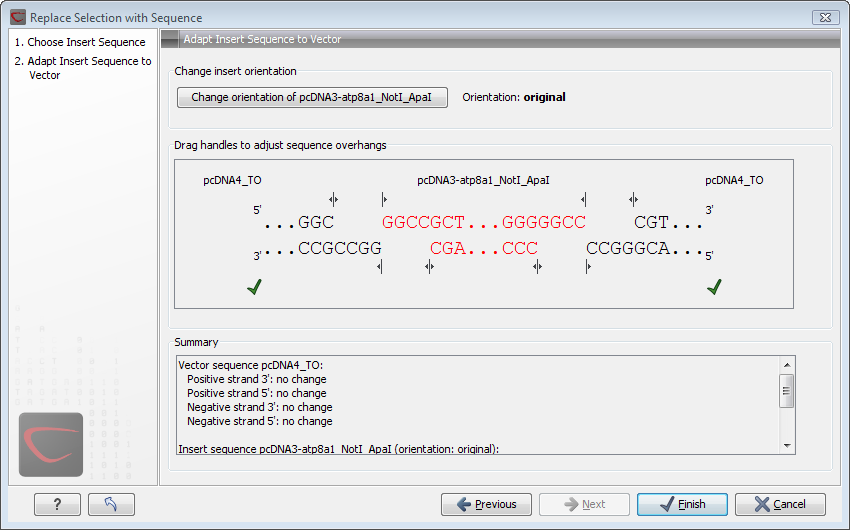
Figure 20.11: Drag the handles to adjust overhangs.
At the top is a button to reverse complement the inserted sequence.
Below is a visualization of the insertion details. The inserted sequence is at the middle shown in red, and the vector has been split at the insertion point and the ends are shown at each side of the inserted sequence.
If the overhangs of the sequence and the vector do not match, you
can blunt end or fill in the overhangs using the drag handles
(![]() ).
).
Whenever you drag the handles, the status of the insertion point is indicated below:
- The overhangs match (
 ).
).
- The overhangs do not match (
 ). In this case,
you will not be able to click Finish. Drag the handles to make
the overhangs match.
). In this case,
you will not be able to click Finish. Drag the handles to make
the overhangs match.
At the bottom of the dialog is a summary field which records all the
changes made to the overhangs. This contents of the summary will
also be written in the history (![]() ) when you click
Finish.
) when you click
Finish.
When you click Finish and the sequence is inserted, it will be marked with a selection.
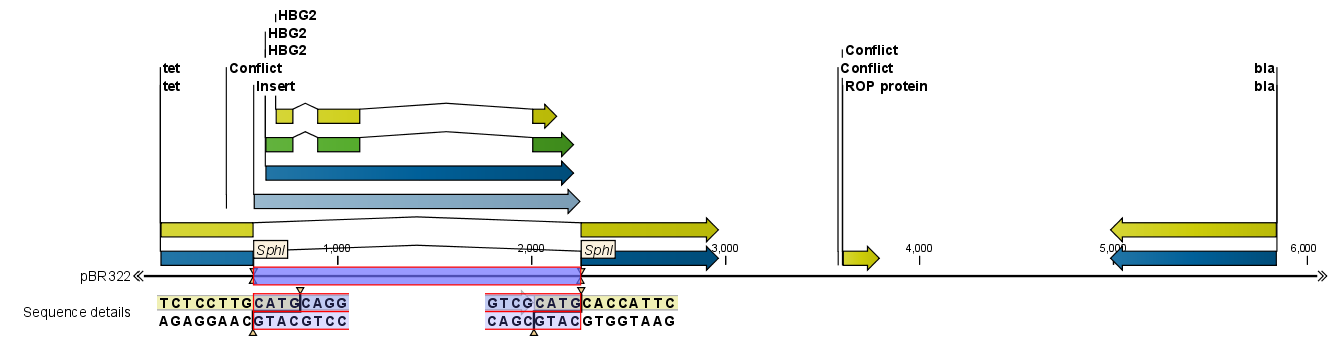
Figure 20.12: One sequence is now inserted into the cloning vector.
The sequence inserted is automatically selected.
