Whole Genome Alignment import and export
Whole Genome Alignments can be exported and imported as MAF and XMFA files using the standard Import and Export functionality of the Workbench (figure 8.1).
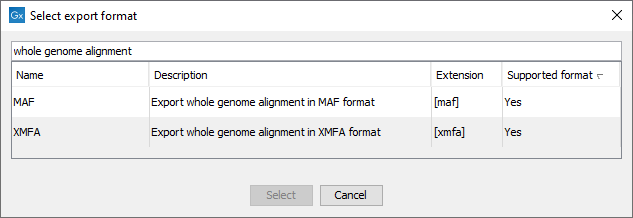
Figure 8.1: Export and import of Whole Genome Alignments can be done using the MAF and XMFA formats.
For MAF and XMFA import, an alignment block must contain each genome only once. We do not support alignment blocks that map multiple times on the same genome.
The MAF importer does not support the optional line types (such as i and e) and these are ignored if found in the file.
The XMFA importer tries to extract the sequence names found as comments in the sequence headers. If the extraction fails, the sequences are named using the numbers found in the file and the comments are ignored.
