Downloading reads and metadata from SRA
Click on Download Reads and Metadata to save reads and their associated data. The data is saved in a metadata table and can be later associated to the reads for use in downstream analysis, for example to define factors for differential expression in the Differential Expression for RNA-Seq tool. Should the metadata table later be deleted, the "Show Metadata for Selection" button can be used to quickly recover a copy without having to re-download all the runs.The Download Reads and Metadata wizard offers the following options:
Import Options
(figure 7.3)
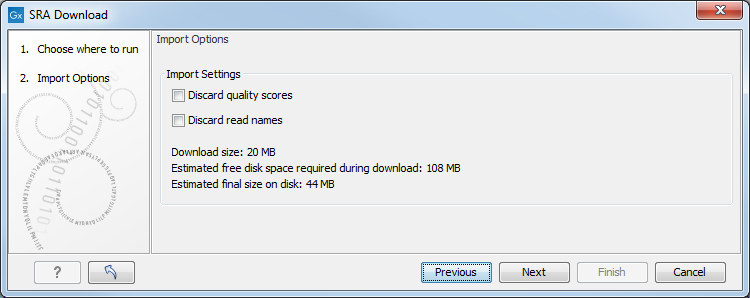
Figure 7.3: The Download Reads and Metadata Import Options dialog.
As with other NGS reads importers, it is possible to discard read names and/or quality scores to save space.
- "Download size" is the size of the .sra files that will be downloaded. Note that in some cases, the actual download may be up to 1GB larger than stated, as .sra files can be reference-compressed, meaning that a copy of the genome must also be retrieved before the file can be converted into fastq and imported into the workbench.
- "Estimated free disk space required during download" is a conservative estimate for the total free disk space required to download the selected runs. This is the "Estimated final size on disk" + the size of the largest single run in FASTQ format + the size of the largest single run in SRA format.
- "Estimated final size on disk" is an estimate of the total size of the files after they have been imported into the workbench.
Edit Paired End Settings
(figure 7.4)
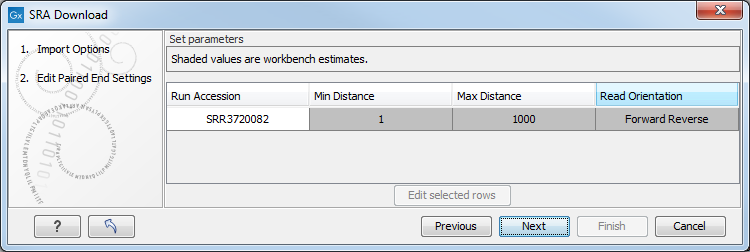
Figure 7.4: The Download Reads and Metadata Edit Paired End Settings dialog.
This dialog appears for all runs marked as being Paired (Paired column contains "Yes").
Read orientation is always guessed to be "Forward Reverse" unless otherwise stated.
Minimum distance and Maximum distance depend on how much data the depositor supplied with the runs. They are allowed to supply an "Insert Size" and an "Insert Deviation".
- If no insert size is supplied, we use defaults of 1 for minimum and 1,000 for maximum.
- If an insert size is supplied, we make the following calculation:
-
- If no deviation is supplied, we estimate this to be
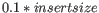 and perform the same calculation as above.
and perform the same calculation as above.
When possible, we generally recommend that SRA data be used in subsequent analyses with the "Auto paired end distance detection" option enabled as the quality of deposited information is low. For example, some depositors report insert size including the length of the reads, and some excluding the length of the reads.
