RNA-Seq and Small RNA analysis
Based on an annotated reference genome, CLC Genomics Workbench supports RNA-Seq Analysis by mapping next-generation sequencing reads and distributing and counting the reads across genes and transcripts. Subsequently, the results can be used for expression analysis. The tools from the RNA-Seq and Small RNA Analysis folder automatically account for differences due to sequencing depth, removing the need to normalize input data.
RNA-Seq analysis, expression analysis, and other tools can be included in workflows. Designing a workflow that includes an RNA-Seq Analysis step, which is typically run once per sample, and an expression analysis step, typically run once to analyze all the samples, is described in Batching part of a workflow.
Metadata and RNA-Seq
The statistical analysis and visualization tools of the RNA-Seq folder make extensive use of the metadata system. For example, metadata are required when defining the experimental design in the Differential Expression for RNA-Seq tool, and can be used to add extra layers of insight in Create Sample Level Heat Map, Create Feature Level Heat Map for RNA-Seq, and PCA for RNA-Seq.
To get the most out of these tools we recommend that all input expression tracks are associated with metadata.
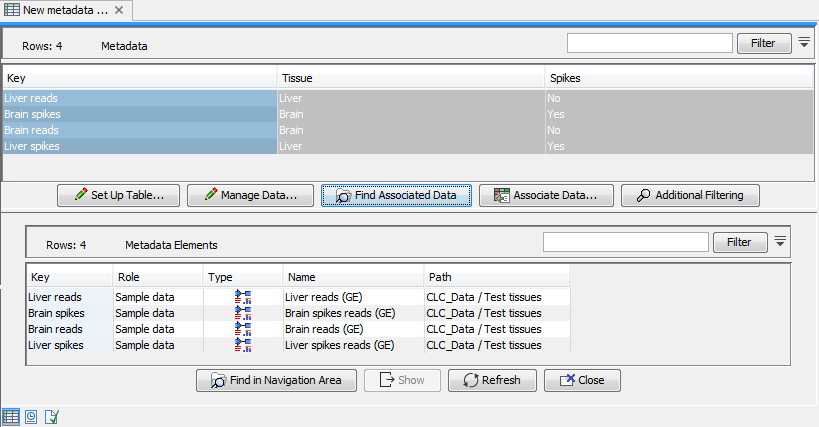
Figure 33.1: A metadata table with expression samples associated with it.
Subsections
- RNA-Seq normalization
- Create Expression Browser
- miRNA analysis
- RNA-Seq Tools
- Expression Plots
- Differential Expression
