Introduction
The Additional Alignments plugin delivers popular, third party alignment tools, complementing the functionality delivered as standard in the CLC software. These are ClustalW, ClustalO, MAFFT and MUSCLE.
After the plugin is installed, these alignment tools are available from the Tools menu (figure 1.1).
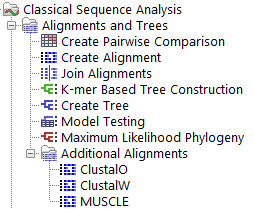
Figure 1.1: The tools provided by this plugin are available from the Additional Alignments folder in the Tools menu.
Tools delivered by this plugin are described briefly in this manual. For more detailed documentation, please refer to the third party websites and research papers.
Note: Annotations on sequences are retained in an alignment, but they are not shown by default. To show sequence annotations when viewing an alignment, select the "Show annotations" option in the Annotation layout palette of the Side Panel.
Subsections
