Introduction
The Ingenuity Pathway Analysis plugin provides the ability to upload expression data from CLC Genomics Workbench to Ingenuity Pathway Analysis (IPA). It is possible to upload:
- Statistical comparison data generated using the RNA-seq tools, and
- Legacy experiment data.
IPA provides valuable biological insight into the results of gene expression experiments by uncovering enriched signaling and metabolic pathways, activated and inhibited upstream regulators and effects on downstream diseases, functions, and phenotypes. IPA can visualize at the isoform level for human genes.
The plugin comes with two tools and a ready-to-use workflow (figure 1.1):
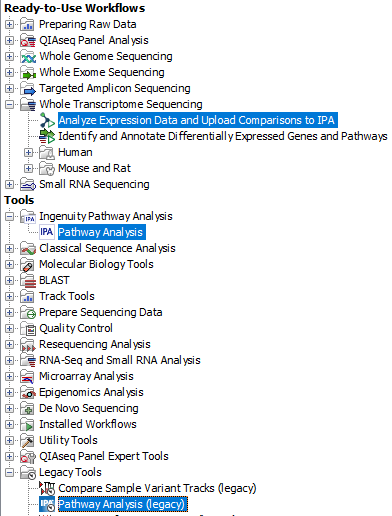
Figure 1.1: IPA plugin tools and workflow when installed in the CLC Genomics Workbench Toolbox.
- The Pathway Analysis tool uploads statistical comparison data (generated by the tool Differential Expression for RNA-Seq) to IPA. The Pathway Analysis tool is based on the legacy Pathway Analysis tool, but has been thoroughly improved on many aspects such as usage of the new IPA API, stability, error handling, and user feedback during the upload process. The tool will output one or more Statistical Comparison tracks.
- The Pathway Analysis (legacy) tool, which can upload legacy experiment data. The tool is primarily included for backwards compatibility reasons.
- The ready-to-use workflow Analyze Expression Data and Upload Comparisons to IPA, which takes expression data as input. The workflow analyzes them using the RNA-Seq Analysis tools, and submits the comparisons to IPA using the Pathway Analysis tool.
It is possible to use gene and transcript based RNA-seq experiments as basis for the analysis, but also microarrays from Illumina and Affymetrix are supported. Currently, small RNA based experiments are not supported by this integration.
Once the experiment data are ready, it is possible to annotate with any of the supported statistics:
- Transformed and normalized foldchange
- Baggerley's test
- Kal's Z test
- ANOVA
- edgeR
