Introduction
The Whole Genome Alignment plugin makes it possible to compare genomes, and to explore and visualize their evolutionary relations (figure 1.1).
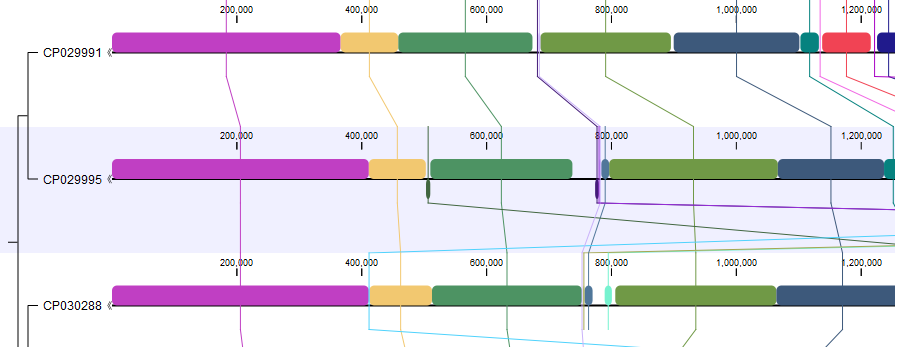
Figure 1.1: A Whole Genome Alignment view.
The plugin installs the tools in a Whole Genome Alignment folder in the Toolbox as seen in figure 1.2, and adds the relevant format files to the Export and Import functionality of the Workbench.

Figure 1.2: The tools are installed in a folder called Whole Genome Alignment in the Toolbox.
The plugin contains functionality for:
- Fast whole genome dot plots
- Aligning multiple genomes
- Visualizing alignments of multiple genomes, which can contain large-scale events such as inversions and translocations
- Calculating the Average Nucleotide Identity
- Transferring annotations from a reference genome to other genomes
- Creating evolutionary trees and heat maps based on the Average Nucleotide Identity
- Importing and exporting standard Whole Genome Alignments formats (MAF and XMFA)
- Extracting multiple sequence alignments, optionally restricting to annotations such as coding regions
Limitations
The tools delivered by this plugin are optimized for working with small and medium-sized genomes (up to 100M bases). They are not designed for larger eukaryotic genomes.
