Predicting Structural Variants
Having created breakpoint signatures (LBs and RBs), we use these in a procedure, which inspects the called breakpoint signatures, and attempts to match and combine them to infer possible underlying structural variant signatures. Based on the inferred structural variant signatures, structural variants are predicted, and annotations created. The procedure for inspecting breakpoint signatures and inferring structural variants is described in detail below.The output of the Structural Variation Plug-in is a 'Structural variants' table and, optionally, a 'Signatures' table and a 'Structural variants report'. The 'Signatures' table contains the LB and RB signature annotations called, and the report gives a summary of the types of unaligned end and structural variant annotations created. When running the analysis on a reads track, the 'Structural variants' and 'Signatures' table outputs are feature tracks. When running the analysis on a stand-alone read mapping, the annotations created are added to the reference sequence of the mapping. The created tables and the mappings are 'linked' allowing the user to easily navigate between the tables and the corresponding positions in the read mapping. Figure 2.3 shows an example of the result of an analysis on a standalone read mapping (to the left) and on a reads track (to the right).
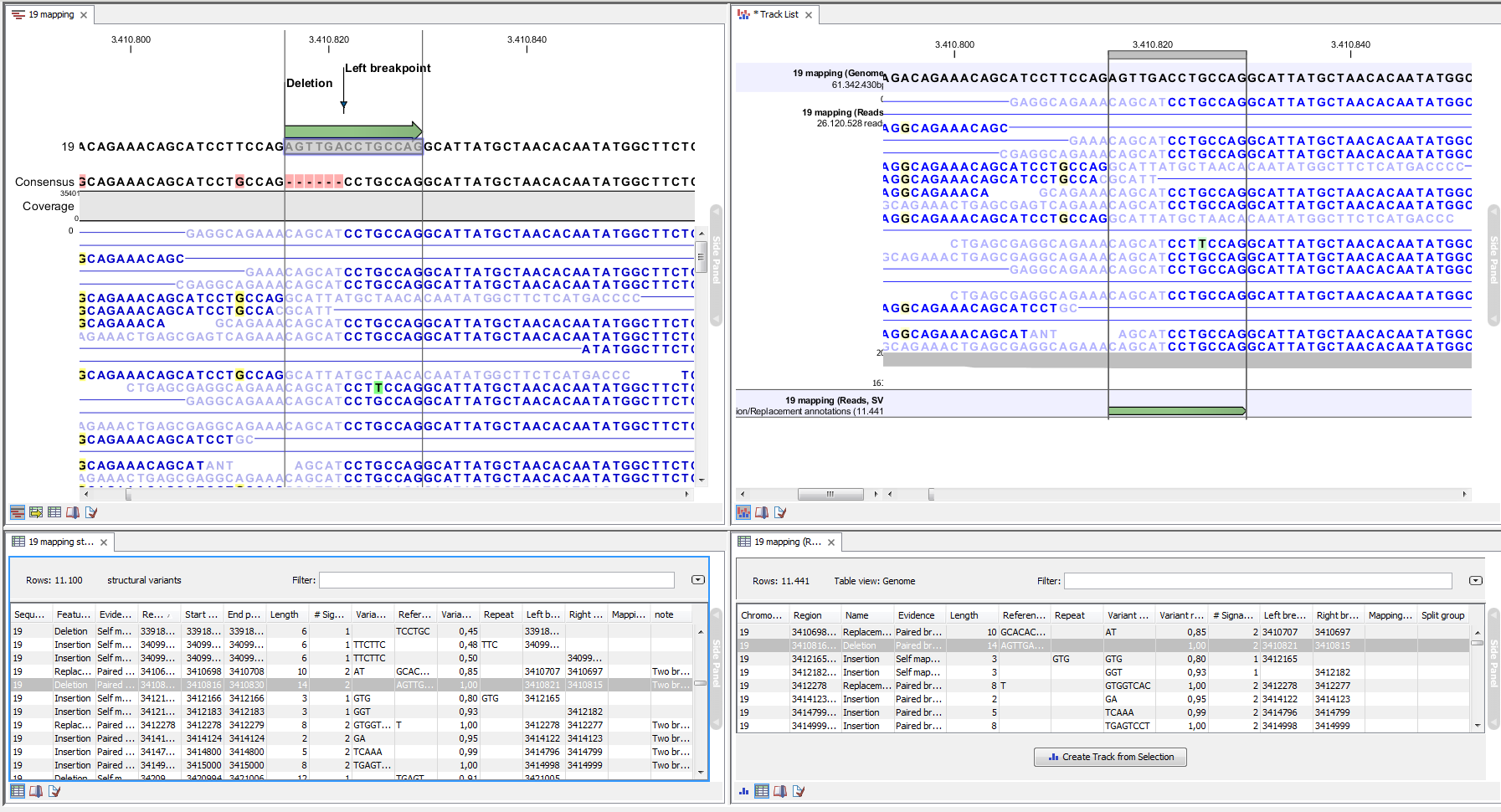
Figure 2.3: Example of the result of an analysis on a standalone read mapping (to the left) and on a reads track (to the right).
Typically, there will be called breakpoint signatures that are not found to stem from a structural variant. There may be a number of reasons for that: (1) the unaligned ends from which the breakpoint signature was derived might not be caused by an underlying structural variant, but merely be due to read mapping issues or noise, or (2) the breakpoint(s) which the detected breakpoint should have been matched to was/were not detected, and therefore no matching breakpoint(s) were found. Breakpoints may go un-detected either because of lack of coverage in the breakpoint region or because they are located within regions with exclusively non-uniquely mapped reads (only unaligned ends of uniquely mapping reads are used).
Subsections
