Signal Peptide Prediction (SignalP 6.0)
The Signal Peptide Prediction (SignalP 6.0) tool uses the SignalP 6.0 service at https://services.healthtech.dtu.dk/services/SignalP-6.0.
The SignalP 6.0 [Teufel et al., 2022] service improves SignalP 4.1 with a new algorithm that use a machine learning model to detect all five SP types and is applicable to metagenomic data. Read more about the differences between Signal 4.1 and 6.0 at https://communities.springernature.com/posts/signalp-6-0-predicts-all-five-types-of-signal-peptides-using-protein-language-models.
The service supports protein sequences with between 5 and 1000 proteins and might timeout if given more than 100 sequences.
To run the Signal Peptide Prediction (SignalP 6) tool, go to:
Toolbox | Classical Sequence Analysis (![]() ) | Protein Analysis (
) | Protein Analysis (![]() ) | Signal Peptide Prediction (SignalP 6.0) (
) | Signal Peptide Prediction (SignalP 6.0) (![]() )
)
In the first wizard step, select a single protein sequence or a protein sequence list (figure 2.4).
Figure 2.4: Select input protein sequences.
Click Next to set the options for the SignalP analysis (figure 2.5).
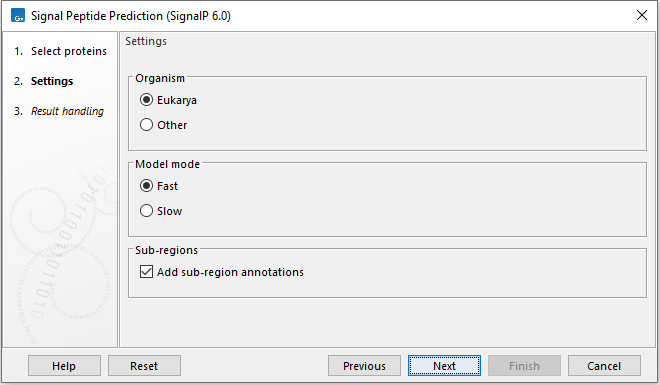
Figure 2.5: Set the options for SignalP 6.0.
- Organism
- Eukarya Use for analyzing Eukarya. When analyzing Eukarya, only "standard" secretory signal peptides transported by the Sec translocon and cleaved by Signal Peptidase I (Lep) are predicted.
- Other Use for analyzing Archaea, Gram-positive Bacteria and Gram-negative Bacteria.
- Model mode
- Fast The analysis mode is fast but region borders may be less accurate.
- Slow The slow mode takes 6x longer to compute. Use when accurate region borders are needed.
- Add sub-region annotatons Add annotations for identified signal peptide sub-regions like n-region, h-region, c-region, cysteine, and twin-arginine motif.
Subsections
