Find Resistance with ShortBRED
This tool allows you to detect and quantify the presence of antibiotic resistance (AR) marker genes by running DIAMOND with a database consisting of peptide marker sequences which represent the genes of interest. These marker databases can be downloaded using the tool Download Resistance Database. The Find Resistance with ShortBRED tool works similarly to the quantify step of ShortBRED, a public bioinformatics pipeline and resource. To learn more about the ShortBRED-Quantify tool [Kaminski et al., 2015], see http://journals.plos.org/ploscompbiol/article?id=10.1371/journal.pcbi.1004557.
Find Resistance with ShortBRED quantifies the presence of Antibiotic Resistance (AR) marker genes in a sample of NGS short reads. It is possible to output a sequence list containing all the input reads which contained a marker (each read in the output is annotated with metadata describing the properties of the marker detected in the read).
To start the tool, go to:
Drug Resistance Analysis (![]() ) | Find Resistance with ShortBRED (
) | Find Resistance with ShortBRED (![]() ).
).
The tool accepts a nucleotide sequence or sequence list as input.
In the next dialog (figure 15.6), several parameters are available:
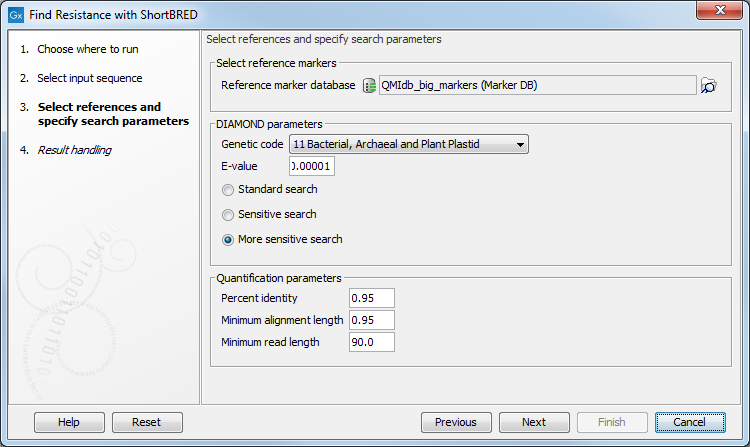
Figure 15.6: References and search parameters.
- Reference sequence database: select the AR marker database to use (downloaded with the tool Download Resistance Database).
- Genetic code: select the genetic code to use when DIAMOND translates the nucleotide sequences to proteins.
- E-value: specify an expectation value to use as threshold for qualifying hits with DIAMOND.
- Standard search: executes DIAMOND in its default mode.
- Sensitive search: executes DIAMOND in its sensitive mode.
- More sensitive search: executes DIAMOND in its most sensitive mode.
- Percent identity: defines a minimum threshold for the percent identity of an alignment. This value is used by Find Resistance with ShortBRED to determine whether a hit found by DIAMOND is sufficiently good to be validated as a true hit (equivalent to the parameter "-id" of the ShortBRED-Quantify tool).
- Minimum alignment length: the minimum length of the DIAMOND alignment. This value is used by Find Resistance with ShortBRED to determine whether a hit found by DIAMOND is sufficiently good to be validated as a true hit (equivalent to the parameter "-pctlength" of the ShortBRED-Quantify tool - see citation at the beginning of this section).
- Minimum read length: the minimum read length. This value is used by Find Resistance with ShortBRED to determine whether a read is long enough to be processed by Find Resistance with ShortBRED (equivalent to the parameter "-minreadBP" of the ShortBRED-Quantify tool).
The Find Resistance with ShortBRED tool will output a resistance abundance table, a result summary table, an optional report, and an optional sequence list.
The result summary table provides an overview of all detected resistance phenotypes. The table reports the number of resistance genes identified, the number of markers and the number of reads for each resistance phenotype.
The optional report output contains general information about the input sample, the marker database used and a short result summary.
The optional sequence list output contains all reads from the input sample which were found to contain one of the AR marker sequences. Each read in the sequence list is annotated with metadata describing the properties of the marker detected in the read.
Subsections
